RUFY3 and RUFY4 are ARL8 effectors that promote coupling of endolysosomes to dynein-dynactin
- PMID: 35314674
- PMCID: PMC8938451
- DOI: 10.1038/s41467-022-28952-y
RUFY3 and RUFY4 are ARL8 effectors that promote coupling of endolysosomes to dynein-dynactin
Abstract
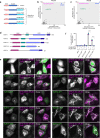
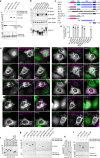
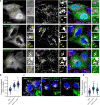
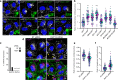
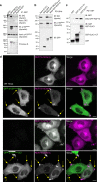
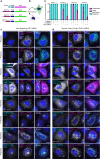
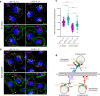
The small GTPase ARL8 associates with endolysosomes, leading to the recruitment of several effectors that couple endolysosomes to kinesins for anterograde transport along microtubules, and to tethering factors for eventual fusion with other organelles. Herein we report the identification of the RUN- and FYVE-domain-containing proteins RUFY3 and RUFY4 as ARL8 effectors that promote coupling of endolysosomes to dynein-dynactin for retrograde transport along microtubules. Using various methodologies, we find that RUFY3 and RUFY4 interact with both GTP-bound ARL8 and dynein-dynactin. In addition, we show that RUFY3 and RUFY4 promote concentration of endolysosomes in the juxtanuclear area of non-neuronal cells, and drive redistribution of endolysosomes from the axon to the soma in hippocampal neurons. The function of RUFY3 in retrograde transport contributes to the juxtanuclear redistribution of endolysosomes upon cytosol alkalinization. These studies thus identify RUFY3 and RUFY4 as ARL8-dependent, dynein-dynactin adaptors or regulators, and highlight the role of ARL8 in the control of both anterograde and retrograde endolysosome transport.
© 2022. This is a U.S. Government work and not under copyright protection in the US; foreign copyright protection may apply.
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
The role of microtubules and the dynein/dynactin motor complex of host cells in the biogenesis of the Coxiella burnetii-containing vacuole.PLoS One. 2019 Jan 14;14(1):e0209820. doi: 10.1371/journal.pone.0209820. eCollection 2019. PLoS One. 2019. PMID: 30640917 Free PMC article.
-
Functional analysis of dynactin and cytoplasmic dynein in slow axonal transport.J Neurosci. 1996 Nov 1;16(21):6742-52. doi: 10.1523/JNEUROSCI.16-21-06742.1996. J Neurosci. 1996. PMID: 8824315 Free PMC article.
-
Ordered recruitment of dynactin to the microtubule plus-end is required for efficient initiation of retrograde axonal transport.J Neurosci. 2013 Aug 7;33(32):13190-203. doi: 10.1523/JNEUROSCI.0935-13.2013. J Neurosci. 2013. PMID: 23926272 Free PMC article.
-
[Dynein and dynactin as organizers of the system of cell microtubules].Ontogenez. 2006 Sep-Oct;37(5):323-39. Ontogenez. 2006. PMID: 17066975 Review. Russian.
-
Choreographing the motor-driven endosomal dance.J Cell Sci. 2023 Mar 1;136(5):jcs259689. doi: 10.1242/jcs.259689. Epub 2022 Nov 16. J Cell Sci. 2023. PMID: 36382597 Free PMC article. Review.
Cited by
-
KIF1C facilitates retrograde transport of lysosomes through Hook3 and dynein.Commun Biol. 2024 Oct 11;7(1):1305. doi: 10.1038/s42003-024-07023-6. Commun Biol. 2024. PMID: 39394274 Free PMC article.
-
A BORC-dependent molecular pathway for vesiculation of cell corpse phagolysosomes.Curr Biol. 2023 Feb 27;33(4):607-621.e7. doi: 10.1016/j.cub.2022.12.041. Epub 2023 Jan 17. Curr Biol. 2023. PMID: 36652947 Free PMC article.
-
Regulation of angiogenesis by endocytic trafficking mediated by cytoplasmic dynein 1 light intermediate chain 1.bioRxiv [Preprint]. 2024 Apr 1:2024.04.01.587559. doi: 10.1101/2024.04.01.587559. bioRxiv. 2024. Update in: Angiogenesis. 2024 Nov;27(4):943-962. doi: 10.1007/s10456-024-09951-7. PMID: 38903077 Free PMC article. Updated. Preprint.
-
Cargo specificity, regulation, and therapeutic potential of cytoplasmic dynein.Exp Mol Med. 2024 Apr;56(4):827-835. doi: 10.1038/s12276-024-01200-7. Epub 2024 Apr 1. Exp Mol Med. 2024. PMID: 38556551 Free PMC article. Review.
-
TMEM55B links autophagy flux, lysosomal repair, and TFE3 activation in response to oxidative stress.Nat Commun. 2024 Jan 2;15(1):93. doi: 10.1038/s41467-023-44316-6. Nat Commun. 2024. PMID: 38168055 Free PMC article.
References
-
- Bagshaw RD, Callahan JW, Mahuran DJ. The Arf-family protein, Arl8b, is involved in the spatial distribution of lysosomes. Biochem. Biophys. Res. Commun. 2006;344:1186–1191. - PubMed
-
- Hofmann I, Munro S. An N-terminally acetylated Arf-like GTPase is localised to lysosomes and affects their motility. J. Cell Sci. 2006;119:1494–1503. - PubMed
-
- Antonny B, Beraud-Dufour S, Chardin P, Chabre M. N-terminal hydrophobic residues of the G-protein ADP-ribosylation factor-1 insert into membrane phospholipids upon GDP to GTP exchange. Biochemistry. 1997;36:4675–4684. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

