Exogenous lipase administration alters gut microbiota composition and ameliorates Alzheimer's disease-like pathology in APP/PS1 mice
- PMID: 35314754
- PMCID: PMC8938460
- DOI: 10.1038/s41598-022-08840-7
Exogenous lipase administration alters gut microbiota composition and ameliorates Alzheimer's disease-like pathology in APP/PS1 mice
Abstract
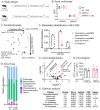
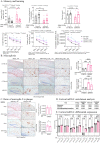
Alzheimer's disease (AD) represents the most common form of dementia in the elderly with no available disease modifying treatments. Altered gut microbial composition has been widely acknowledged as a common feature of AD, which potentially contributes to progression or onset of AD. To assess the hypothesis that Candida rugosa lipase (CRL), which has been shown to enhance gut microbiome and metabolite composition, can rebalance the gut microbiome composition and reduce AD pathology, the treatment effects in APPswe/PS1de9 (APP/PS1) mice were investigated. The analysis revealed an increased abundance of Acetatifactor and Clostridiales vadin BB60 genera in the gut; increased lipid hydrolysis in the gut lumen, normalization of peripheral unsaturated fatty acids, and reduction of neuroinflammation and memory deficits post treatment. Finally, we demonstrated that the evoked benefits on memory could be transferred via fecal matter transplant (FMT) into antibiotic-induced microbiome-depleted (AIMD) wildtype mice, ameliorating their memory deficits. The findings herein contributed to improve our understanding of the role of the gut microbiome in AD's complex networks and suggested that targeted modification of the gut could contribute to amelioration of AD neuropathology.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Alzheimer's Disease Histological and Behavioral Manifestations in Transgenic Mice Correlate with Specific Gut Microbiome State.J Alzheimers Dis. 2017;56(1):385-390. doi: 10.3233/JAD-160884. J Alzheimers Dis. 2017. PMID: 27911317
-
Human gut microbiota Agathobaculum butyriciproducens improves cognitive impairment in LPS-induced and APP/PS1 mouse models of Alzheimer's disease.Nutr Res. 2021 Feb;86:96-108. doi: 10.1016/j.nutres.2020.12.010. Epub 2020 Dec 9. Nutr Res. 2021. PMID: 33551257
-
Jatrorrhizine Balances the Gut Microbiota and Reverses Learning and Memory Deficits in APP/PS1 transgenic mice.Sci Rep. 2019 Dec 20;9(1):19575. doi: 10.1038/s41598-019-56149-9. Sci Rep. 2019. PMID: 31862965 Free PMC article.
-
Role of gut-brain axis, gut microbial composition, and probiotic intervention in Alzheimer's disease.Life Sci. 2021 Jan 1;264:118627. doi: 10.1016/j.lfs.2020.118627. Epub 2020 Oct 22. Life Sci. 2021. PMID: 33169684 Review.
-
The Effect of Gut Microbe Dysbiosis on the Pathogenesis of Alzheimer's Disease (AD) and Related Conditions.Curr Alzheimer Res. 2022;19(4):274-284. doi: 10.2174/1567205019666220419101205. Curr Alzheimer Res. 2022. PMID: 35440296 Review.
Cited by
-
The Effects of a Microbial Enzyme Mixture on Macronutrient Hydrolysis in a Static Simulation of Oro-Gastric Digestion That Models Human Digestive Senescence.Foods. 2025 Mar 10;14(6):937. doi: 10.3390/foods14060937. Foods. 2025. PMID: 40231923 Free PMC article.
-
Gut microbiota-host lipid crosstalk in Alzheimer's disease: implications for disease progression and therapeutics.Mol Neurodegener. 2024 Apr 16;19(1):35. doi: 10.1186/s13024-024-00720-0. Mol Neurodegener. 2024. PMID: 38627829 Free PMC article. Review.
-
Potential Roles of Exogenous Proteases and Lipases as Prebiotics.Nutrients. 2025 Mar 6;17(5):924. doi: 10.3390/nu17050924. Nutrients. 2025. PMID: 40077794 Free PMC article. Review.
-
Causal associations between gut microbiota, metabolites, and idiopathic normal pressure hydrocephalus: a two‑sample Mendelian randomization study.BMC Neurol. 2025 Jun 9;25(1):249. doi: 10.1186/s12883-025-04187-4. BMC Neurol. 2025. PMID: 40490712 Free PMC article.
-
Nutrient-Synbiotic Complex Ameliorates LPS-Induced Depressive-Like Behavior via Modulation of Gut Microbiota and Tryptophan Metabolism in Mice.Food Sci Nutr. 2025 Jul 15;13(7):e70628. doi: 10.1002/fsn3.70628. eCollection 2025 Jul. Food Sci Nutr. 2025. PMID: 40672547 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

