Single-cell transcriptome analysis reveals three sequential phases of gene expression during zebrafish sensory hair cell regeneration
- PMID: 35316618
- PMCID: PMC9188816
- DOI: 10.1016/j.devcel.2022.03.001
Single-cell transcriptome analysis reveals three sequential phases of gene expression during zebrafish sensory hair cell regeneration
Abstract
Loss of sensory hair cells (HCs) in the mammalian inner ear leads to permanent hearing and vestibular defects, whereas loss of HCs in zebrafish results in their regeneration. We used single-cell RNA sequencing (scRNA-seq) to characterize the transcriptional dynamics of HC regeneration in zebrafish at unprecedented spatiotemporal resolution. We uncovered three sequentially activated modules: first, an injury/inflammatory response and downregulation of progenitor cell maintenance genes within minutes after HC loss; second, the transient activation of regeneration-specific genes; and third, a robust re-activation of developmental gene programs, including HC specification, cell-cycle activation, ribosome biogenesis, and a metabolic switch to oxidative phosphorylation. The results are relevant not only for our understanding of HC regeneration and how we might be able to trigger it in mammals but also for regenerative processes in general. The data are searchable and publicly accessible via a web-based interface.
Keywords: hair cell death; hair cell regeneration; lateral line system; mechanosensory organ; neomycin; pseudotime analysis; scRNA-seq atlas; stem cells; support cells; zebrafish.
Copyright © 2022 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures






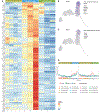


References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

