Factors regulating cellulolytic gene expression in filamentous fungi: an overview
- PMID: 35317826
- PMCID: PMC8939176
- DOI: 10.1186/s12934-022-01764-x
Factors regulating cellulolytic gene expression in filamentous fungi: an overview
Abstract
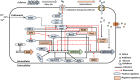
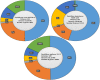
The growing demand for biofuels such as bioethanol has led to the need for identifying alternative feedstock instead of conventional substrates like molasses, etc. Lignocellulosic biomass is a relatively inexpensive feedstock that is available in abundance, however, its conversion to bioethanol involves a multistep process with different unit operations such as size reduction, pretreatment, saccharification, fermentation, distillation, etc. The saccharification or enzymatic hydrolysis of cellulose to glucose involves a complex family of enzymes called cellulases that are usually fungal in origin. Cellulose hydrolysis requires the synergistic action of several classes of enzymes, and achieving the optimum secretion of these simultaneously remains a challenge. The expression of fungal cellulases is controlled by an intricate network of transcription factors and sugar transporters. Several genetic engineering efforts have been undertaken to modulate the expression of cellulolytic genes, as well as their regulators. This review, therefore, focuses on the molecular mechanism of action of these transcription factors and their effect on the expression of cellulases and hemicellulases.
Keywords: Catabolite repression; Cellulase expression; Regulation; Transcription factors; Transporters.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
Similar articles
-
Cellulolytic enzyme production and enzymatic hydrolysis for second-generation bioethanol production.Adv Biochem Eng Biotechnol. 2012;128:1-24. doi: 10.1007/10_2011_131. Adv Biochem Eng Biotechnol. 2012. PMID: 22231654 Review.
-
Bioconversion of lignocellulosic biomass: biochemical and molecular perspectives.J Ind Microbiol Biotechnol. 2008 May;35(5):377-391. doi: 10.1007/s10295-008-0327-8. Epub 2008 Mar 13. J Ind Microbiol Biotechnol. 2008. PMID: 18338189 Review.
-
Synergism of fungal and bacterial cellulases and hemicellulases: a novel perspective for enhanced bio-ethanol production.Biotechnol Lett. 2015 Jun;37(6):1117-29. doi: 10.1007/s10529-015-1779-3. Epub 2015 Feb 6. Biotechnol Lett. 2015. PMID: 25656474 Review.
-
Addressing challenges in production of cellulases for biomass hydrolysis: Targeted interventions into the genetics of cellulase producing fungi.Bioresour Technol. 2021 Jun;329:124746. doi: 10.1016/j.biortech.2021.124746. Epub 2021 Jan 29. Bioresour Technol. 2021. PMID: 33610429 Review.
-
Engineering Robust Cellulases for Tailored Lignocellulosic Degradation Cocktails.Int J Mol Sci. 2020 Feb 26;21(5):1589. doi: 10.3390/ijms21051589. Int J Mol Sci. 2020. PMID: 32111065 Free PMC article. Review.
Cited by
-
Correlating sugar transporter expression and activities to identify transporters for an orphan sugar substrate.Appl Microbiol Biotechnol. 2024 Dec;108(1):83. doi: 10.1007/s00253-023-12907-4. Epub 2024 Jan 8. Appl Microbiol Biotechnol. 2024. PMID: 38189952 Free PMC article.
-
Dual Role of MtHAC-1 in Regulating Cellulase and Xylanase Production in Myceliophthora thermophila.Microb Biotechnol. 2025 Aug;18(8):e70203. doi: 10.1111/1751-7915.70203. Microb Biotechnol. 2025. PMID: 40739661 Free PMC article.
-
Structural and biochemical insights of xylose MFS and SWEET transporters in microbial cell factories: challenges to lignocellulosic hydrolysates fermentation.Front Microbiol. 2024 Sep 27;15:1452240. doi: 10.3389/fmicb.2024.1452240. eCollection 2024. Front Microbiol. 2024. PMID: 39397797 Free PMC article. Review.
-
Genomic Characterization and Establishment of a Genetic Manipulation System for Trichoderma sp. (Harzianum Clade) LZ117.J Fungi (Basel). 2024 Oct 7;10(10):697. doi: 10.3390/jof10100697. J Fungi (Basel). 2024. PMID: 39452649 Free PMC article.
-
Different Putative Methyltransferases Have Different Effects on the Expression Patterns of Cellulolytic Genes.J Fungi (Basel). 2023 Nov 17;9(11):1118. doi: 10.3390/jof9111118. J Fungi (Basel). 2023. PMID: 37998923 Free PMC article.
References
-
- Schifter I, Diaz L, Rodriguez R, Gomez JP, Gonzalez U. Combustion and emissions behavior for ethanol–gasoline blends in a single cylinder engine. Fuels. 2011;90:3586–3592.
-
- Singh M, Shukla R, Das K. Harvesting of microalgal biomass. In: Bux F, editor. Biotechnological applications of microalgae: biodiesel and value added products. Taylor and Francis; 2013. pp. 75–87.
-
- Huber GW, Dale BE. Grassoline at the pump. Sci Am. 2009;301:52–59. - PubMed
-
- Chandel AK, Singh OV, Venkateswar Rao L, Chandrasekhar G, Lakshmi NM. Bioconversion of novel substrate Saccharum spontaneum, a weedy material, into ethanol by Pichia stipitis NCIM3498. Bioresource Technol. 2011;102:1709–1714. - PubMed
-
- Limayem A, Ricke SC. Lignocellulosic biomass for bioethanol production: current perspectives, potential issues and future prospects. Prog Energ Combust. 2012;38:449–467.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

