Structural perspectives on the mechanism of signal activation, ligand selectivity and allosteric modulation in angiotensin receptors: IUPHAR Review 34
- PMID: 35318654
- PMCID: PMC9398925
- DOI: 10.1111/bph.15840
Structural perspectives on the mechanism of signal activation, ligand selectivity and allosteric modulation in angiotensin receptors: IUPHAR Review 34
Abstract
Functional advances have guided our knowledge of physiological and fatal pathological mechanisms of the hormone angiotensin II (AngII) and its antagonists. Such studies revealed that tissue response to a given dose of the hormone or its antagonist depends on receptors that engage the ligand. Thus, we need to know much more about the structures of receptor-ligand complexes at high resolution. Recently, X-ray structures of both AngII receptors (AT1 and AT2 receptors) bound to peptide and non-peptide ligands have been elucidated, providing new opportunities to examine the dynamic fluxes in the 3D architecture of the receptors, as the basis of ligand selectivity, efficacy, and regulation of the molecular functions of the receptors. Constituent structural motifs cooperatively transform ligand selectivity into specific functions, thus conceptualizing the primacy of the 3D structure over individual motifs of receptors. This review covers the new data elucidating the structural dynamics of AngII receptors and how structural knowledge can be transformative in understanding the mechanisms underlying the physiology of AngII.
Keywords: AT1 receptor; AT2 receptor; AngII; G protein; GPCR; allosteric modulation; autoantibody; ligand selectivity; β-arrestin.
© 2022 The British Pharmacological Society.
Conflict of interest statement
Conflict of Interest
The authors have no conflicts of interest to declare.
Figures
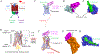
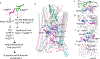
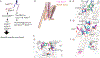
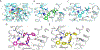
Similar articles
-
The arrestin-selective angiotensin AT1 receptor agonist [Sar1,Ile4,Ile8]-AngII negatively regulates bradykinin B2 receptor signaling via AT1-B2 receptor heterodimers.J Biol Chem. 2013 Jun 28;288(26):18872-84. doi: 10.1074/jbc.M113.472381. Epub 2013 May 9. J Biol Chem. 2013. PMID: 23661707 Free PMC article.
-
Angiotensin II cyclic analogs as tools to investigate AT1R biased signaling mechanisms.Biochem Pharmacol. 2018 Aug;154:104-117. doi: 10.1016/j.bcp.2018.04.021. Epub 2018 Apr 20. Biochem Pharmacol. 2018. PMID: 29684376
-
Differential beta-arrestin binding of AT1 and AT2 angiotensin receptors.FEBS Lett. 2006 Jan 9;580(1):41-5. doi: 10.1016/j.febslet.2005.11.044. Epub 2005 Dec 6. FEBS Lett. 2006. PMID: 16359671
-
Computational approaches on angiotensin receptors and their ligands: recent developments and results.Curr Med Chem. 2007;14(29):3105-21. doi: 10.2174/092986707782793970. Curr Med Chem. 2007. PMID: 18220745 Review.
-
Angiotensin II receptors.J Am Soc Nephrol. 1999 Jan;10 Suppl 11:S30-9. J Am Soc Nephrol. 1999. PMID: 9892138 Review.
Cited by
-
Analysis of peri-implant bone tissue between hydrophilic and rough implant surfaces in spontaneously hypertensive rats treated with losartan.J Appl Oral Sci. 2024 Jun 24;32:e20230374. doi: 10.1590/1678-7757-2023-0374. eCollection 2024. J Appl Oral Sci. 2024. PMID: 38922240 Free PMC article.
-
The Angiotensin AT2 Receptor: From a Binding Site to a Novel Therapeutic Target.Pharmacol Rev. 2022 Oct;74(4):1051-1135. doi: 10.1124/pharmrev.120.000281. Pharmacol Rev. 2022. PMID: 36180112 Free PMC article. Review.
-
Allosteric Regulation of G-Protein-Coupled Receptors: From Diversity of Molecular Mechanisms to Multiple Allosteric Sites and Their Ligands.Int J Mol Sci. 2023 Mar 24;24(7):6187. doi: 10.3390/ijms24076187. Int J Mol Sci. 2023. PMID: 37047169 Free PMC article. Review.
-
Gating Mechanism for Biased Agonism at Angiotensin II Type 1 Receptors.Molecules. 2025 May 30;30(11):2399. doi: 10.3390/molecules30112399. Molecules. 2025. PMID: 40509287 Free PMC article.
References
-
- Allen AM, Dosanjh JK, Erac M, Dassanayake S, Hannan RD, & Thomas WG (2006). Expression of constitutively active angiotensin receptors in the rostral ventrolateral medulla increases blood pressure. Hypertension 47: 1054–1061. - PubMed
-
- Alexander W, Bernstein KE, Catt KJ, de Gasparo M, Dhanachandra Singh K, Eguchi, et al. (2019) “Angiotensin receptors (version 2019.4) in the IUPHAR/BPS Guide to Pharmacology Database”, IUPHAR/BPS Guide to Pharmacology CITE, 2019(4). doi: 10.2218/gtopdb/F6/2019.4. - DOI
-
- Andreas S, Herrmann-Lingen C, Raupach T, Luthje L, Fabricius JA, Hruska N, et al. (2006). Angiotensin II blockers in obstructive pulmonary disease: a randomised controlled trial. Eur Respir J 27: 972–979. - PubMed
-
- Asada H, Horita S, Hirata K, Shiroishi M, Shiimura Y, Iwanari H, et al. (2018). Crystal structure of the human angiotensin II type 2 receptor bound to an angiotensin II analog. Nat Struct Mol Biol 25: 570–576. - PubMed
-
- Asada H, Inoue A, Ngako Kadji FM, Hirata K, Shiimura Y, Im D, et al. (2020). The Crystal Structure of Angiotensin II Type 2 Receptor with Endogenous Peptide Hormone. Structure 28: 418–425 e414. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

