Rhes protein transits from neuron to neuron and facilitates mutant huntingtin spreading in the brain
- PMID: 35319973
- PMCID: PMC8942366
- DOI: 10.1126/sciadv.abm3877
Rhes protein transits from neuron to neuron and facilitates mutant huntingtin spreading in the brain
Abstract
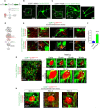
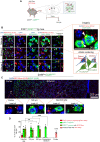
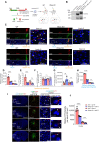
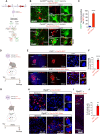
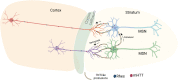
Rhes (RASD2) is a thyroid hormone-induced gene that regulates striatal motor activity and promotes neurodegeneration in Huntington disease (HD) and tauopathy. Rhes moves and transports the HD protein, polyglutamine-expanded huntingtin (mHTT), via tunneling nanotube (TNT)-like membranous protrusions between cultured neurons. However, similar intercellular Rhes transportation in the intact brain was unknown. Here, we report that Rhes induces TNT-like protrusions in the striatal medium spiny neurons (MSNs) and transported between dopamine-1 receptor (D1R)-MSNs and D2R-MSNs of intact striatum and organotypic brain slices. Notably, mHTT is robustly transported within the striatum and from the striatum to the cortical areas in the brain, and Rhes deletion diminishes such transport. Moreover, Rhes moves to the cortical regions following restricted expression in the MSNs of the striatum. Thus, Rhes is a first striatum-enriched protein demonstrated to move and transport mHTT between neurons and brain regions, providing new insights into interneuronal protein transport in the brain.
Figures
Similar articles
-
Striatal Induction and Spread of the Huntington's Disease Protein: A Novel Rhes Route.J Huntingtons Dis. 2022;11(3):281-290. doi: 10.3233/JHD-220548. J Huntingtons Dis. 2022. PMID: 35871361 Free PMC article. Review.
-
Ectopic expression of the striatal-enriched GTPase Rhes elicits cerebellar degeneration and an ataxia phenotype in Huntington's disease.Neurobiol Dis. 2015 Oct;82:66-77. doi: 10.1016/j.nbd.2015.05.011. Epub 2015 Jun 3. Neurobiol Dis. 2015. PMID: 26048156
-
Rhes travels from cell to cell and transports Huntington disease protein via TNT-like protrusion.J Cell Biol. 2019 Jun 3;218(6):1972-1993. doi: 10.1083/jcb.201807068. Epub 2019 May 10. J Cell Biol. 2019. PMID: 31076452 Free PMC article.
-
The role of Rhes, Ras homolog enriched in striatum, in neurodegenerative processes.Exp Cell Res. 2013 Sep 10;319(15):2310-5. doi: 10.1016/j.yexcr.2013.03.033. Epub 2013 Apr 10. Exp Cell Res. 2013. PMID: 23583659 Review.
-
Bioinformatics analysis of Ras homologue enriched in the striatum, a potential target for Huntington's disease therapy.Int J Mol Med. 2019 Dec;44(6):2223-2233. doi: 10.3892/ijmm.2019.4373. Epub 2019 Oct 15. Int J Mol Med. 2019. PMID: 31638189 Free PMC article.
Cited by
-
Striatal Induction and Spread of the Huntington's Disease Protein: A Novel Rhes Route.J Huntingtons Dis. 2022;11(3):281-290. doi: 10.3233/JHD-220548. J Huntingtons Dis. 2022. PMID: 35871361 Free PMC article. Review.
-
Selective deletion of zinc transporter 3 in amacrine cells promotes retinal ganglion cell survival and optic nerve regeneration after injury.Neural Regen Res. 2023 Dec;18(12):2773-2780. doi: 10.4103/1673-5374.373660. Neural Regen Res. 2023. PMID: 37449644 Free PMC article.
-
Astroglia proliferate upon the biogenesis of tunneling nanotubes via α-synuclein dependent transient nuclear translocation of focal adhesion kinase.iScience. 2024 Jul 23;27(8):110565. doi: 10.1016/j.isci.2024.110565. eCollection 2024 Aug 16. iScience. 2024. PMID: 39184442 Free PMC article.
-
Role of SUMOylation in Neurodegenerative Diseases.Cells. 2022 Oct 27;11(21):3395. doi: 10.3390/cells11213395. Cells. 2022. PMID: 36359791 Free PMC article. Review.
-
Invisible Bridges: Unveiling the Role and Prospects of Tunneling Nanotubes in Cancer Therapy.Mol Pharm. 2024 Nov 4;21(11):5413-5429. doi: 10.1021/acs.molpharmaceut.4c00563. Epub 2024 Oct 7. Mol Pharm. 2024. PMID: 39373242 Free PMC article. Review.
References
-
- Vargiu P., Morte B., Manzano J., Perez J., de Abajo R., Gregor Sutcliffe J., Bernal J., Thyroid hormone regulation of rhes, a novel Ras homolog gene expressed in the striatum. Brain Res. Mol. Brain Res. 94, 1–8 (2001). - PubMed
-
- Falk J. D., Vargiu P., Foye P. E., Usui H., Perez J., Danielson P. E., Lerner D. L., Bernal J., Sutcliffe J. G., Rhes: A striatal-specific Ras homolog related to Dexras1. J. Neurosci. Res. 57, 782–788 (1999). - PubMed
-
- Napolitano F., D’Angelo L., de Girolamo P., Avallone L., de Lange P., Usiello A., The thyroid hormone-target gene rhes a novel crossroad for neurological and psychiatric disorders: New insights from animal models. Neuroscience 384, 419–428 (2018). - PubMed
-
- Shahani N., Swarnkar S., Giovinazzo V., Morgenweck J., Bohn L. M., Scharager-Tapia C., Pascal B., Martinez-Acedo P., Khare K., Subramaniam S., RasGRP1 promotes amphetamine-induced motor behavior through a Rhes interaction network (“Rhesactome”) in the striatum. Sci. Signal. 9, ra111 (2016). - PMC - PubMed
-
- Napolitano F., De Rosa A., Russo R., Di Maio A., Garofalo M., Federici M., Migliarini S., Ledonne A., Rizzo F. R., Avallone L., Nuzzo T., Biagini T., Pasqualetti M., Mercuri N. B., Mazza T., Chambery A., Usiello A., The striatal-enriched protein Rhes is a critical modulator of cocaine-induced molecular and behavioral responses. Sci. Rep. 9, 15294 (2019). - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

