Human papillomavirus type 16 E6 induces cell competition
- PMID: 35320322
- PMCID: PMC8979454
- DOI: 10.1371/journal.ppat.1010431
Human papillomavirus type 16 E6 induces cell competition
Abstract
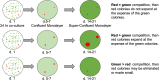
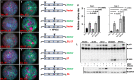
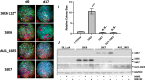
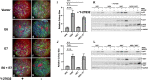
High-risk human papillomavirus (HPV) infections induce squamous epithelial tumors in which the virus replicates. Initially, the virus-infected cells are untransformed, but expand in both number and area at the expense of uninfected squamous epithelial cells. We have developed an in vitro assay in which colonies of post-confluent HPV16 expressing cells outcompete and displace confluent surrounding uninfected keratinocytes. The enhanced colony competition induced by the complete HPV16 genome is conferred by E6 expression alone, not by individual expression of E5 or E7, and requires E6 interaction with p53. E6-expressing keratinocytes undermine and displace adjacent normal keratinocytes from contact with the attachment substrate, thereby expanding the area of the E6-expressing colony at the expense of normal keratinocytes. These new results separate classic oncogenicity that is primarily conferred by HPV16 E7 from cell competition that we show is primarily conferred by E6 and provides a new biological role for E6 oncoproteins from high-risk human papillomaviruses.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
ELF3 Overexpression Contributes to the Malignant Transformation of HPV16 E6/E7-Immortalized Keratinocytes by Promoting CCNE2 Expression.J Microbiol Biotechnol. 2024 Dec 28;34(12):2484-2491. doi: 10.4014/jmb.2408.08041. Epub 2024 Oct 30. J Microbiol Biotechnol. 2024. PMID: 39572025 Free PMC article.
-
Functional interaction between human papillomavirus type 16 E6 and E7 oncoproteins and cigarette smoke components in lung epithelial cells.PLoS One. 2012;7(5):e38178. doi: 10.1371/journal.pone.0038178. Epub 2012 May 25. PLoS One. 2012. PMID: 22662279 Free PMC article.
-
Human Papillomavirus E6/E7 and Long Noncoding RNA TMPOP2 Mutually Upregulated Gene Expression in Cervical Cancer Cells.J Virol. 2019 Apr 3;93(8):e01808-18. doi: 10.1128/JVI.01808-18. Print 2019 Apr 15. J Virol. 2019. PMID: 30728257 Free PMC article.
-
Manipulation of Epithelial Differentiation by HPV Oncoproteins.Viruses. 2019 Apr 22;11(4):369. doi: 10.3390/v11040369. Viruses. 2019. PMID: 31013597 Free PMC article. Review.
-
High-Risk Human Papillomavirus Oncogenic E6/E7 mRNAs Splicing Regulation.Front Cell Infect Microbiol. 2022 Jun 27;12:929666. doi: 10.3389/fcimb.2022.929666. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 35832386 Free PMC article. Review.
Cited by
-
HPV E6 inhibits E6AP to regulate epithelial homeostasis by modulating keratinocyte differentiation commitment and YAP1 activation.PLoS Pathog. 2023 Jun 28;19(6):e1011464. doi: 10.1371/journal.ppat.1011464. eCollection 2023 Jun. PLoS Pathog. 2023. PMID: 37379354 Free PMC article.
-
Human papillomavirus associated cervical lesion: pathogenesis and therapeutic interventions.MedComm (2020). 2023 Sep 14;4(5):e368. doi: 10.1002/mco2.368. eCollection 2023 Oct. MedComm (2020). 2023. PMID: 37719443 Free PMC article. Review.
-
Conditional Cell Reprogramming and Air-Liquid Interface Modeling Life Cycle of Oncogenic Viruses (HPV and EBV) in Epithelial Cells and Virus-Associated Human Carcinomas.Viruses. 2023 Jun 17;15(6):1388. doi: 10.3390/v15061388. Viruses. 2023. PMID: 37376685 Free PMC article. Review.
-
How can HPV E6 manipulate host cell differentiation process to maintain the reservoir of infection.Tumour Virus Res. 2025 Jun;19:200313. doi: 10.1016/j.tvr.2025.200313. Epub 2025 Jan 19. Tumour Virus Res. 2025. PMID: 39832674 Free PMC article. No abstract available.
-
Human papillomaviruses: Knowns, mysteries, and unchartered territories.J Med Virol. 2023 Oct;95(10):e29191. doi: 10.1002/jmv.29191. J Med Virol. 2023. PMID: 37861365 Free PMC article. Review.
References
-
- Knipe D, Howley PM. Virology. Lippincott Williams and Wilkins; 2007. p. 1662.
-
- Brandsma JL, Yang ZH, Barthold SW, Johnson EA. Use of a rapid, efficient inoculation method to induce papillomas by cottontail rabbit papillomavirus DNA shows that the E7 gene is required. Proc Natl Acad Sci U S A. 1991;88(11):4816–20. doi: 10.1073/pnas.88.11.4816 ; PubMed Central PMCID: PMC51757. - DOI - PMC - PubMed
-
- Wu X, Xiao W, Brandsma JL. Papilloma formation by cottontail rabbit papillomavirus requires E1 and E2 regulatory genes in addition to E6 and E7 transforming genes. J Virol. 1994;68(9):6097–102. Epub 1994/09/01. doi: 10.1128/JVI.68.9.6097-6102.1994 ; PubMed Central PMCID: PMC237021. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

