Differential Gain-of-Function Activity of Three p53 Hotspot Mutants In Vivo
- PMID: 35320355
- PMCID: PMC9117479
- DOI: 10.1158/0008-5472.CAN-21-3376
Differential Gain-of-Function Activity of Three p53 Hotspot Mutants In Vivo
Abstract
The majority of TP53 missense mutations identified in cancer patients are in the DNA-binding domain and are characterized as either structural or contact mutations. These missense mutations exhibit inhibitory effects on wild-type p53 activity. More importantly, these mutations also demonstrate gain-of-function (GOF) activities characterized by increased metastasis, poor prognosis, and drug resistance. To better understand the activities by which TP53 mutations, identified in Li-Fraumeni syndrome, contribute to tumorigenesis, we generated mice harboring a novel germline Trp53R245W allele (contact mutation) and compared them with existing models with Trp53R172H (structural mutation) and Trp53R270H (contact mutation) alleles. Thymocytes from heterozygous mice showed that all three hotspot mutations exhibited similar inhibitory effects on wild-type p53 transcription in vivo, and tumors from these mice had similar levels of loss of heterozygosity. However, the overall survival of Trp53R245W/+ and Trp53R270H/+ mice, but not Trp53R172H/+ mice, was significantly shorter than that of Trp53+/- mice, providing strong evidence for p53-mutant-specific GOF contributions to tumor development. Furthermore, Trp53R245W/+ and Trp53R270H/+ mice had more osteosarcoma metastases than Trp53R172H/+ mice, suggesting that these two contact mutants have stronger GOF in driving osteosarcoma metastasis. Transcriptomic analyses using RNA sequencing data from Trp53R172H/+, Trp53R245W/+, and Trp53R270H/+ primary osteosarcomas in comparison with Trp53+/- indicated that GOF of the three mutants was mediated by distinct pathways. Thus, both the inhibitory effect of mutant over wild-type p53 and GOF activities of mutant p53 contributed to tumorigenesis in vivo. Targeting p53 mutant-specific pathways may be important for therapeutic outcomes in osteosarcoma.
Significance: p53 hotspot mutants inhibit wild-type p53 similarly but differ in their GOF activities, with stronger tumor-promoting activity in contact mutants and distinct protein partners of each mutant driving tumorigenesis and metastasis.
©2022 American Association for Cancer Research.
Conflict of interest statement
Competing interests
The authors declare no competing interests.
The authors declare no potential conflicts of interest
Figures

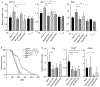



References
-
- Patch AM, Christie EL, Etemadmoghadam D, Garsed DW, George J, Fereday S, et al.. Whole-genome characterization of chemoresistant ovarian cancer. Nature 2015;521:489–94 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

