eIF6 rebinding dynamically couples ribosome maturation and translation
- PMID: 35322020
- PMCID: PMC8943182
- DOI: 10.1038/s41467-022-29214-7
eIF6 rebinding dynamically couples ribosome maturation and translation
Abstract
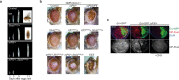
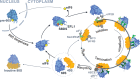
Protein synthesis is a cyclical process consisting of translation initiation, elongation, termination and ribosome recycling. The release factors SBDS and EFL1-both mutated in the leukemia predisposition disorder Shwachman-Diamond syndrome - license entry of nascent 60S ribosomal subunits into active translation by evicting the anti-association factor eIF6 from the 60S intersubunit face. We find that in mammalian cells, eIF6 holds all free cytoplasmic 60S subunits in a translationally inactive state and that SBDS and EFL1 are the minimal components required to recycle these 60S subunits back into additional rounds of translation by evicting eIF6. Increasing the dose of eIF6 in mice in vivo impairs terminal erythropoiesis by sequestering post-termination 60S subunits in the cytoplasm, disrupting subunit joining and attenuating global protein synthesis. These data reveal that ribosome maturation and recycling are dynamically coupled by a mechanism that is disrupted in an inherited leukemia predisposition disorder.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





Similar articles
-
Uncoupling of GTP hydrolysis from eIF6 release on the ribosome causes Shwachman-Diamond syndrome.Genes Dev. 2011 May 1;25(9):917-29. doi: 10.1101/gad.623011. Genes Dev. 2011. PMID: 21536732 Free PMC article.
-
Mechanism of eIF6 release from the nascent 60S ribosomal subunit.Nat Struct Mol Biol. 2015 Nov;22(11):914-9. doi: 10.1038/nsmb.3112. Epub 2015 Oct 19. Nat Struct Mol Biol. 2015. PMID: 26479198 Free PMC article.
-
Somatic genetic rescue of a germline ribosome assembly defect.Nat Commun. 2021 Aug 19;12(1):5044. doi: 10.1038/s41467-021-24999-5. Nat Commun. 2021. PMID: 34413298 Free PMC article.
-
eIF6 anti-association activity is required for ribosome biogenesis, translational control and tumor progression.Biochim Biophys Acta. 2015 Jul;1849(7):830-5. doi: 10.1016/j.bbagrm.2014.09.010. Epub 2014 Sep 22. Biochim Biophys Acta. 2015. PMID: 25252159 Review.
-
Molecular basis of the human ribosomopathy Shwachman-Diamond syndrome.Adv Biol Regul. 2018 Jan;67:109-127. doi: 10.1016/j.jbior.2017.09.002. Epub 2017 Sep 6. Adv Biol Regul. 2018. PMID: 28942353 Free PMC article. Review.
Cited by
-
Predisposition to myeloid malignancies in Shwachman-Diamond syndrome: biological insights and clinical advances.Blood. 2023 Mar 30;141(13):1513-1523. doi: 10.1182/blood.2022017739. Blood. 2023. PMID: 36542827 Free PMC article. Review.
-
Shwachman-Diamond syndromes: clinical, genetic, and biochemical insights from the rare variants.Haematologica. 2023 Oct 1;108(10):2594-2605. doi: 10.3324/haematol.2023.282949. Haematologica. 2023. PMID: 37226705 Free PMC article.
-
Sequestration of ribosomal subunits as inactive 80S by targeting eIF6 limits mitotic exit and cancer progression.Nucleic Acids Res. 2025 Feb 8;53(4):gkae1272. doi: 10.1093/nar/gkae1272. Nucleic Acids Res. 2025. PMID: 39727167 Free PMC article.
-
On the origin of the nucleus: a hypothesis.Microbiol Mol Biol Rev. 2023 Dec 20;87(4):e0018621. doi: 10.1128/mmbr.00186-21. Epub 2023 Nov 29. Microbiol Mol Biol Rev. 2023. PMID: 38018971 Free PMC article. Review.
-
CLPP-Null Eukaryotes with Excess Heme Biosynthesis Show Reduced L-arginine Levels, Probably via CLPX-Mediated OAT Activation.Biomolecules. 2024 Feb 19;14(2):241. doi: 10.3390/biom14020241. Biomolecules. 2024. PMID: 38397478 Free PMC article.
References
-
- Ceci M, et al. Release of eIF6 (p27BBP) from the 60S subunit allows 80S ribosome assembly. Nature. 2003;426:579–584. - PubMed
-
- Russell DW, Spremulli LL. Mechanism of action of the wheat germ ribosome dissociation factor: interaction with the 60 S subunit. Arch. Biochem. Biophys. 1980;201:518–526. - PubMed
-
- Senger B, et al. The nucle(ol)ar Tif6p and Efl1p are required for a late cytoplasmic step of ribosome synthesis. Mol. Cell. 2001;8:1363–1373. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

