Single cell transcriptional diversity and intercellular crosstalk of human liver cancer
- PMID: 35322024
- PMCID: PMC8943132
- DOI: 10.1038/s41419-022-04689-w
Single cell transcriptional diversity and intercellular crosstalk of human liver cancer
Abstract
Liver cancer arises from the evolutionary selection of the dynamic tumor microenvironment (TME), in which the tumor cell generally becomes more heterogeneous; however, the mechanisms of TME-mediated transcriptional diversity of liver cancer remain unclear. Here, we assess transcriptional diversity in 15 liver cancer patients by single-cell transcriptome analysis and observe transcriptional diversity of tumor cells is associated with stemness in liver cancer patients. Tumor-associated fibroblast (TAF), as a potential driving force behind the heterogeneity in tumor cells within and between tumors, was predicted to interact with high heterogeneous tumor cells via COL1A1-ITGA2. Moreover, COL1A1-mediated YAP-signaling activation might be the mechanistic link between TAF and tumor cells with increased transcriptional diversity. Strikingly, the levels of COL1A1, ITGA2, and YAP are associated with morphological heterogeneity and poor overall survival of liver cancer patients. Beyond providing a potential mechanistic link between the TME and heterogeneous tumor cells, this study establishes that collagen-stimulated YAP activation is associates with transcriptional diversity in tumor cells by upregulating stemness, providing a theoretical basis for individualized treatment targets.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

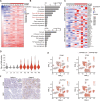



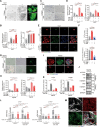
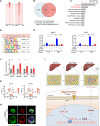
Similar articles
-
Single-cell transcriptomic architecture and intercellular crosstalk of human intrahepatic cholangiocarcinoma.J Hepatol. 2020 Nov;73(5):1118-1130. doi: 10.1016/j.jhep.2020.05.039. Epub 2020 Jun 5. J Hepatol. 2020. PMID: 32505533
-
Cancer-Associated Fibroblast-Mediated Cellular Crosstalk Supports Hepatocellular Carcinoma Progression.Hepatology. 2021 May;73(5):1717-1735. doi: 10.1002/hep.31792. Hepatology. 2021. PMID: 33682185
-
A Dynamic Transcriptome Map of Different Tissue Microenvironment Cells Identified During Gastric Cancer Development Using Single-Cell RNA Sequencing.Front Immunol. 2021 Oct 21;12:728169. doi: 10.3389/fimmu.2021.728169. eCollection 2021. Front Immunol. 2021. PMID: 34745098 Free PMC article.
-
The Role of Cancer-Associated Fibroblasts and Fibrosis in Liver Cancer.Annu Rev Pathol. 2017 Jan 24;12:153-186. doi: 10.1146/annurev-pathol-052016-100322. Epub 2016 Dec 5. Annu Rev Pathol. 2017. PMID: 27959632 Free PMC article. Review.
-
Significance of the Tumor Microenvironment in Liver Cancer Progression.Crit Rev Oncog. 2020;25(1):1-9. doi: 10.1615/CritRevOncog.2020034987. Crit Rev Oncog. 2020. PMID: 32865906 Review.
Cited by
-
Advances in single-cell RNA sequencing and its applications in cancer research.J Hematol Oncol. 2023 Aug 24;16(1):98. doi: 10.1186/s13045-023-01494-6. J Hematol Oncol. 2023. PMID: 37612741 Free PMC article. Review.
-
Cancer-Associated Fibroblasts in Hepatocellular Carcinoma and Cholangiocarcinoma.Cell Mol Gastroenterol Hepatol. 2023;15(4):985-999. doi: 10.1016/j.jcmgh.2023.01.006. Epub 2023 Jan 25. Cell Mol Gastroenterol Hepatol. 2023. PMID: 36708970 Free PMC article. Review.
-
Crosstalk between 5-methylcytosine and N6-methyladenosine machinery defines disease progression, therapeutic response and pharmacogenomic landscape in hepatocellular carcinoma.Mol Cancer. 2023 Jan 10;22(1):5. doi: 10.1186/s12943-022-01706-6. Mol Cancer. 2023. PMID: 36627693 Free PMC article.
-
Fetal liver development and implications for liver disease pathogenesis.Nat Rev Gastroenterol Hepatol. 2023 Sep;20(9):561-581. doi: 10.1038/s41575-023-00775-2. Epub 2023 May 19. Nat Rev Gastroenterol Hepatol. 2023. PMID: 37208503 Review.
-
Integrin α2 in the microenvironment and the tumor compartment of digestive (gastrointestinal) cancers: emerging regulators and therapeutic opportunities.Front Oncol. 2024 Nov 6;14:1439709. doi: 10.3389/fonc.2024.1439709. eCollection 2024. Front Oncol. 2024. PMID: 39568561 Free PMC article. Review.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

