Localization and functional characterization of the alternative oxidase in Naegleria
- PMID: 35322502
- PMCID: PMC9540462
- DOI: 10.1111/jeu.12908
Localization and functional characterization of the alternative oxidase in Naegleria
Abstract
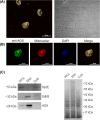
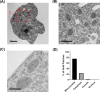
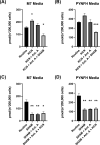
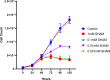
The alternative oxidase (AOX) is a protein involved in supporting enzymatic reactions of the Krebs cycle in instances when the canonical (cytochrome-mediated) respiratory chain has been inhibited, while allowing for the maintenance of cell growth and necessary metabolic processes for survival. Among eukaryotes, alternative oxidases have dispersed distribution and are found in plants, fungi, and protists, including Naegleria ssp. Naegleria species are free-living unicellular amoeboflagellates and include the pathogenic species of N. fowleri, the so-called "brain-eating amoeba." Using a multidisciplinary approach, we aimed to understand the evolution, localization, and function of AOX and the role that plays in Naegleria's biology. Our analyses suggest that AOX was present in last common ancestor of the genus and structure prediction showed that all functional residues are also present in Naegleria species. Using cellular and biochemical techniques, we also functionally characterize N. gruberi's AOX in its mitochondria, and we demonstrate that its inactivation affects its proliferation. Consequently, we discuss the benefits of the presence of this protein in Naegleria species, along with its potential pathogenicity role in N. fowleri. We predict that our findings will spearhead new explorations to understand the cell biology, metabolism, and evolution of Naegleria and other free-living relatives.
Keywords: Naegleria; adaptation; confocal microscopy; evolution; mitochondria; respiration.
© 2022 The Authors. Journal of Eukaryotic Microbiology published by Wiley Periodicals LLC on behalf of International Society of Protistologists.
Figures


Similar articles
-
Naegleria's mitotic spindles are built from unique tubulins and highlight core spindle features.Curr Biol. 2022 Mar 28;32(6):1247-1261.e6. doi: 10.1016/j.cub.2022.01.034. Epub 2022 Feb 8. Curr Biol. 2022. PMID: 35139359 Free PMC article.
-
The genome of Naegleria gruberi illuminates early eukaryotic versatility.Cell. 2010 Mar 5;140(5):631-42. doi: 10.1016/j.cell.2010.01.032. Cell. 2010. PMID: 20211133
-
The genome of Naegleria lovaniensis, the basis for a comparative approach to unravel pathogenicity factors of the human pathogenic amoeba N. fowleri.BMC Genomics. 2018 Sep 5;19(1):654. doi: 10.1186/s12864-018-4994-1. BMC Genomics. 2018. PMID: 30185166 Free PMC article.
-
Alternative oxidase: distribution, induction, properties, structure, regulation, and functions.Biochemistry (Mosc). 2014 Dec;79(13):1615-34. doi: 10.1134/S0006297914130112. Biochemistry (Mosc). 2014. PMID: 25749168 Review.
-
[Alternative oxidase - never ending story].Postepy Biochem. 2016;62(2):138-148. Postepy Biochem. 2016. PMID: 28132465 Review. Polish.
Cited by
-
Copper Metabolism in Naegleria gruberi and Its Deadly Relative Naegleria fowleri.Front Cell Dev Biol. 2022 Apr 11;10:853463. doi: 10.3389/fcell.2022.853463. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 35478954 Free PMC article.
References
-
- Barsottini, M.R.O. , Copsey, A. , Young, L. , Baroni, R.M. , Cordeiro, A.T. , Pereira, G.A.G. et al. (2020) Biochemical characterization and inhibition of the alternative oxidase enzyme from the fungal phytopathogen moniliophthora perniciosa. Communications Biology, 3, 263. Available from: 10.1038/s42003-020-0981-6 - DOI - PMC - PubMed
-
- Bexkens, M.L. , Zimorski, V. , Sarink, M.J. , Wienk, H. , Brouwers, J.F. , De Jonckheere, J.F. et al. (2018) Lipids are the preferred substrate of the protist Naegleria gruberi, relative of a human brain pathogen. Cell Reports, 25(3), 537–543.e3. Available from: 10.1016/j.celrep.2018.09.055 - DOI - PMC - PubMed
-
- Chalmers, R.M. (2014) Chapter twenty – Naegleria . In: Percival, S.L. , Yates, M.V. , Williams, D.W. , Chalmers, R.M. & Gray, N.F. (Eds.) Microbiology of waterborne diseases, 2nd Edition. London: Academic Press, pp. 407–416. Available from: 10.1016/B978-0-12-415846-7.00020-2 - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

