Twelve Variants Polygenic Score for Low-Density Lipoprotein Cholesterol Distribution in a Large Cohort of Patients With Clinically Diagnosed Familial Hypercholesterolemia With or Without Causative Mutations
- PMID: 35322671
- PMCID: PMC9075429
- DOI: 10.1161/JAHA.121.023668
Twelve Variants Polygenic Score for Low-Density Lipoprotein Cholesterol Distribution in a Large Cohort of Patients With Clinically Diagnosed Familial Hypercholesterolemia With or Without Causative Mutations
Abstract
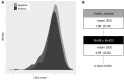
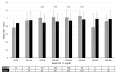
Background A significant proportion of individuals clinically diagnosed with familial hypercholesterolemia (FH), but without any disease-causing mutation, are likely to have polygenic hypercholesterolemia. We evaluated the distribution of a polygenic risk score, consisting of 12 low-density lipoprotein cholesterol (LDL-C)-raising variants (polygenic LDL-C risk score), in subjects with a clinical diagnosis of FH. Methods and Results Within the Lipid Transport Disorders Italian Genetic Network (LIPIGEN) study, 875 patients who were FH-mutation positive (women, 54.75%; mean age, 42.47±15.00 years) and 644 patients who were FH-mutation negative (women, 54.21%; mean age, 49.73±13.54 years) were evaluated. Patients who were FH-mutation negative had lower mean levels of pretreatment LDL-C than patients who were FH-mutation positive (217.14±55.49 versus 270.52±68.59 mg/dL, P<0.0001). The mean value (±SD) of the polygenic LDL-C risk score was 1.00 (±0.18) in patients who were FH-mutation negative and 0.94 (±0.20) in patients who were FH-mutation positive (P<0.0001). In the receiver operating characteristic analysis, the area under the curve for recognizing subjects characterized by polygenic hypercholesterolemia was 0.59 (95% CI, 0.56-0.62), with sensitivity and specificity being 78% and 36%, respectively, at 0.905 as a cutoff value. Higher mean polygenic LDL-C risk score levels were observed among patients who were FH-mutation negative having pretreatment LDL-C levels in the range of 150 to 350 mg/dL (150-249 mg/dL: 1.01 versus 0.91, P<0.0001; 250-349 mg/dL: 1.02 versus 0.95, P=0.0001). A positive correlation between polygenic LDL-C risk score and pretreatment LDL-C levels was observed among patients with FH independently of the presence of causative mutations. Conclusions This analysis confirms the role of polymorphisms in modulating LDL-C levels, even in patients with genetically confirmed FH. More data are needed to support the use of the polygenic score in routine clinical practice.
Keywords: familial hypercholesterolemia; molecular diagnosis; polygenic risk score.
Figures

References
-
- Zamora A, Masana L, Comas‐Cufi M, Vila A, Plana N, Garcia‐Gil M, Alves‐Cabratosa L, Marrugat J, Roman I, Ramos R, et al. Familial hypercholesterolemia in a European Mediterranean population‐Prevalence and clinical data from 2.5 million primary care patients. J Clin Lipidol. 2017;11:1013–1022. doi: 10.1016/j.jacl.2017.05.012 - DOI - PubMed
-
- Nordestgaard BG, Chapman MJ, Humphries SE, Ginsberg HN, Masana L, Descamps OS, Wiklund O, Hegele RA, Raal FJ, Defesche JC, et al. Familial hypercholesterolaemia is underdiagnosed and undertreated in the general population: guidance for clinicians to prevent coronary heart disease: consensus statement of the European Atherosclerosis Society. Eur Heart J. 2013;34:3478–3490. doi: 10.1093/eurheartj/eht273 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

