CHROMOMETHYLTRANSFERASE3/KRYPTONITE maintains the sulfurea paramutation in Solanum lycopersicum
- PMID: 35324329
- PMCID: PMC9060480
- DOI: 10.1073/pnas.2112240119
CHROMOMETHYLTRANSFERASE3/KRYPTONITE maintains the sulfurea paramutation in Solanum lycopersicum
Abstract
SignificanceParamutation involves the transfer of a repressive epigenetic mark between silent and active alleles. It is best known from exceptional non-Mendelian inheritance of conspicuous phenotypes in maize but also in other plants and animals. Recent genomic studies, however, indicate that paramutation may be less exceptional. It may be a consequence of wide-cross hybridization and may contribute to quantitative trait variation or unstable phenotypes in crops. Using the sulfurea (sulf) locus in tomato, we demonstrate that a self-reinforcing feedback loop involving DNA- and histone-methyl transferases CHROMOMETHYLTRANSFERASE3 (CMT3) and KRYPTONITE (KYP) is required for paramutation of sulf and that there is a change in chromatin organization. These findings advance the understanding of non-Mendelian inheritance in plants.
Keywords: epigenetic memory; heredity; paramutation; tomato.
Conflict of interest statement
The authors declare no competing interest.
Figures



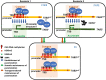
References
-
- Hollick J. B., Paramutation and related phenomena in diverse species. Nat. Rev. Genet. 18, 5–23 (2017). - PubMed
-
- Pilu R., Paramutation phenomena in plants. Semin. Cell Dev. Biol. 44, 2–10 (2015). - PubMed
-
- Hövel I., Pearson N. A., Stam M., Cis-acting determinants of paramutation. Semin. Cell Dev. Biol. 44, 22–32 (2015). - PubMed
-
- Ellew C. P., The genetics of “rogues” among culinary peas (Pisum sativum). Proc. R. Soc. London. Ser. B, Contain. Pap. a Biol. Character 91, 186–195 (1920).
-
- Hagemann R., [Somatic conversion in Lycopersicon esculentum Mill] [Article in German]. Z. Vererbungsl. 89, 587–613 (1958). - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

