Novel Antimicrobial Peptides Designed Using a Recurrent Neural Network Reduce Mortality in Experimental Sepsis
- PMID: 35326874
- PMCID: PMC8944797
- DOI: 10.3390/antibiotics11030411
Novel Antimicrobial Peptides Designed Using a Recurrent Neural Network Reduce Mortality in Experimental Sepsis
Abstract
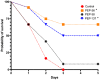
The search and development of new antibiotics is relevant due to widespread antibiotic resistance. One of the promising strategies is the de novo design of novel antimicrobial peptides. The amino acid sequences of 198 novel peptides were obtained using a generative long short-term memory recurrent neural network (LSTM RNN). To assess their antimicrobial effect, we synthesized five out of 198 generated peptides. The PEP-38 and PEP-137 peptides were active in vitro against carbapenem-resistant isolates of Klebsiella aerogenes and K. pneumoniae. PEP-137 was also active against Pseudomonas aeruginosa. The remaining three peptides (PEP-36, PEP-136 and PEP-174) showed no antibacterial effect. Then the effect of PEP-38 and PEP-137 (a single intraperitoneal administration of a 100 μg dose 30 min after infection) on animal survival in an experimental murine model of K. pneumoniae-induced sepsis was investigated. As a control, two groups of mice were used: one received sterile saline, and the other received inactive in vitro PEP-36 (a single 100 μg dose). The PEP-36 peptide was shown to provide the highest survival rate (66.7%). PEP-137 showed a survival rate of 50%. PEP-38 was found to be ineffective. The data obtained can be used to develop new antibacterial peptide drugs to combat antibiotic resistance.
Keywords: LSTM; RNN; antibiotic resistance; antimicrobial peptide; carbapenem-resistance; neural network; peptide design; sepsis.
Conflict of interest statement
The author declares no conflict of interest. The funder had no role in the design of the study, in the collection, analyses, or interpretation of data, in the writing of the manuscript, or in the decision to publish the results.
Figures
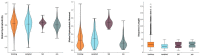

Similar articles
-
Antimicrobial Peptides Epinecidin-1 and Beta-Defesin-3 Are Effective against a Broad Spectrum of Antibiotic-Resistant Bacterial Isolates and Increase Survival Rate in Experimental Sepsis.Antibiotics (Basel). 2022 Jan 9;11(1):76. doi: 10.3390/antibiotics11010076. Antibiotics (Basel). 2022. PMID: 35052952 Free PMC article.
-
Computational antimicrobial peptide design and evaluation against multidrug-resistant clinical isolates of bacteria.J Biol Chem. 2018 Mar 9;293(10):3492-3509. doi: 10.1074/jbc.M117.805499. Epub 2017 Dec 19. J Biol Chem. 2018. PMID: 29259134 Free PMC article.
-
Recurrent Neural Network Model for Constructive Peptide Design.J Chem Inf Model. 2018 Feb 26;58(2):472-479. doi: 10.1021/acs.jcim.7b00414. Epub 2018 Jan 22. J Chem Inf Model. 2018. PMID: 29355319
-
Cost-effectiveness of antimicrobial treatment for inpatients with carbapenem-resistant Klebsiella pneumoniae infection: a systematic review of economic evidence.JBI Database System Rev Implement Rep. 2019 Dec;17(12):2417-2451. doi: 10.11124/JBISRIR-D-18-00019. JBI Database System Rev Implement Rep. 2019. PMID: 31821188
-
De novo designed lipopolysaccharide binding peptides: structure based development of antiendotoxic and antimicrobial drugs.Curr Med Chem. 2010;17(27):3080-93. doi: 10.2174/092986710791959756. Curr Med Chem. 2010. PMID: 20629624 Review.
Cited by
-
Andrographolide and 4-Phenylbutyric Acid Administration Increase the Expression of Antimicrobial Peptides Beta-Defensin-1 and Cathelicidin and Reduce Mortality in Murine Sepsis.Antibiotics (Basel). 2022 Nov 15;11(11):1629. doi: 10.3390/antibiotics11111629. Antibiotics (Basel). 2022. PMID: 36421273 Free PMC article.
-
Designing Novel Antimicrobial Agents from the Synthetic Antimicrobial Peptide (Pep-38) to Combat Antibiotic Resistance.Pharmaceuticals (Basel). 2025 Jun 10;18(6):862. doi: 10.3390/ph18060862. Pharmaceuticals (Basel). 2025. PMID: 40573257 Free PMC article.
-
Antibiotics-free compounds for managing carbapenem-resistant bacteria; a narrative review.Front Pharmacol. 2024 Sep 17;15:1467086. doi: 10.3389/fphar.2024.1467086. eCollection 2024. Front Pharmacol. 2024. PMID: 39355778 Free PMC article. Review.
-
Accelerating the Discovery and Design of Antimicrobial Peptides with Artificial Intelligence.Methods Mol Biol. 2024;2714:329-352. doi: 10.1007/978-1-0716-3441-7_18. Methods Mol Biol. 2024. PMID: 37676607
-
A Systematic Review of Deep Learning Methodologies Used in the Drug Discovery Process with Emphasis on In Vivo Validation.Int J Mol Sci. 2023 Mar 31;24(7):6573. doi: 10.3390/ijms24076573. Int J Mol Sci. 2023. PMID: 37047543 Free PMC article.
References
-
- Baturin V., Shchetinin E., Demidenko I., Kunitsina E., Korableva O., Baturina M., Kharitonova Y. Evaluation of Klebsiella Spp. and Acinetobacter Spp. Antibiotic Resistance in Hospital Environment (Stavropol, Russia) Med. News North Cauc. 2014;9 doi: 10.14300/MNNC.2014.09052. - DOI
-
- Minaev S.V., Filipeva N.V., Leskin V.V., Shchetinin E.V., Golubeva M.V., Rakitina E.N., Shamadaev E.Z., Zhdanova T.V. Microbiological Spectrum of Pyoinflammatory Diseases Causative Agents in Children at a Multi-Speciality Hospital. Med. News North Cauc. 2018;13:112–114. doi: 10.14300/MNNC.2018.13032. - DOI
-
- Langford B.J., So M., Raybardhan S., Leung V., Westwood D., MacFadden D.R., Soucy J.-P.R., Daneman N. Bacterial Co-Infection and Secondary Infection in Patients with COVID-19: A Living Rapid Review and Meta-Analysis. Clin. Microbiol. Infect. 2020;26:1622–1629. doi: 10.1016/j.cmi.2020.07.016. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

