ER-phagy in the Occurrence and Development of Cancer
- PMID: 35327508
- PMCID: PMC8945671
- DOI: 10.3390/biomedicines10030707
ER-phagy in the Occurrence and Development of Cancer
Abstract
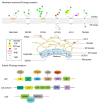
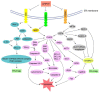
As an organelle, the endoplasmic reticulum (ER) is closely related to protein synthesis and modification. When physiological or pathological stimuli induce disorders of ER function, misfolded proteins trigger ER-phagy, which is beneficial for restoring cell homeostasis or promoting cell apoptosis. As a double-edged sword, ER-phagy actively participates in various stages of development and progression in tumor cells, regulating tumorigenesis and maintaining tumor cell homeostasis. Through the unfolded protein response (UPR), the B cell lymphoma 2 (BCL-2) protein family, the Caspase signaling pathway, and others, ER-phagy plays an initiating role in tumor occurrence, migration, stemness, and proliferation. At the same time, many vital proteins strongly associated with ER-phagy, such as family with sequence similarity 134 member B (FAM134B), translocation protein SEC62 (SEC62), and C/EBP-homologous protein (CHOP), can produce a marked effect in many complex environments, which ultimately lead to entirely different tumor fates. Our article comprehensively focused on introducing the relationship and interaction between ER-phagy and cancers, as well as their molecular mechanism and regulatory pathways. Via these analyses, we tried to clarify the possibility of ER-phagy as a potential target for cancer therapy and provide ideas for further research.
Keywords: ER-phagy; apoptosis; cancer; unfolded protein response.
Conflict of interest statement
The authors declare that there are no conflict of interest to disclose.
Figures
Similar articles
-
FAM134B-induced endoplasmic reticulum (ER)-phagy exacerbates cisplatin-insulted hair cell apoptosis :Possible relation to excessive ER stress.Arch Biochem Biophys. 2023 Oct 15;748:109766. doi: 10.1016/j.abb.2023.109766. Epub 2023 Oct 9. Arch Biochem Biophys. 2023. PMID: 37813237
-
Excessive ER-phagy mediated by the autophagy receptor FAM134B results in ER stress, the unfolded protein response, and cell death in HeLa cells.J Biol Chem. 2019 Dec 27;294(52):20009-20023. doi: 10.1074/jbc.RA119.008709. Epub 2019 Nov 20. J Biol Chem. 2019. PMID: 31748416 Free PMC article.
-
Family with sequence similarity 134 member B-mediated reticulophagy ameliorates hepatocyte apoptosis induced by dithiothreitol.World J Gastroenterol. 2022 Jun 21;28(23):2569-2581. doi: 10.3748/wjg.v28.i23.2569. World J Gastroenterol. 2022. PMID: 35949353 Free PMC article.
-
ER stress as a trigger of UPR and ER-phagy in cancer growth and spread.Front Oncol. 2022 Nov 3;12:997235. doi: 10.3389/fonc.2022.997235. eCollection 2022. Front Oncol. 2022. PMID: 36408145 Free PMC article.
-
Crosstalk of ER stress-mediated autophagy and ER-phagy: Involvement of UPR and the core autophagy machinery.J Cell Physiol. 2018 May;233(5):3867-3874. doi: 10.1002/jcp.26137. Epub 2017 Aug 30. J Cell Physiol. 2018. PMID: 28777470 Review.
Cited by
-
Selective autophagy in cancer: mechanisms, therapeutic implications, and future perspectives.Mol Cancer. 2024 Jan 24;23(1):22. doi: 10.1186/s12943-024-01934-y. Mol Cancer. 2024. PMID: 38262996 Free PMC article. Review.
-
Integrative Analysis of Radiation-Induced Senescence-Associated Secretory Phenotype Factors in Kidney Cancer Progression.Genes (Basel). 2025 Jan 15;16(1):85. doi: 10.3390/genes16010085. Genes (Basel). 2025. PMID: 39858632 Free PMC article.
-
Application of calcium overload-based ion interference therapy in tumor treatment: strategies, outcomes, and prospects.Front Pharmacol. 2024 Feb 15;15:1352377. doi: 10.3389/fphar.2024.1352377. eCollection 2024. Front Pharmacol. 2024. PMID: 38425645 Free PMC article. Review.
-
Human Blood Serum Inhibits Ductal Carcinoma Cells BT474 Growth and Modulates Effect of HER2 Inhibition.Biomedicines. 2022 Aug 8;10(8):1914. doi: 10.3390/biomedicines10081914. Biomedicines. 2022. PMID: 36009461 Free PMC article.
-
Membrane Contact Sites in Proteostasis and ER Stress Response.Contact (Thousand Oaks). 2025 Jul 28;8:25152564251363050. doi: 10.1177/25152564251363050. eCollection 2025 Jan-Dec. Contact (Thousand Oaks). 2025. PMID: 40735229 Free PMC article. Review.
References
Publication types
Grants and funding
- 2016YFA0201504/National Key Research and Development Program of China
- 81473249 and 81102464/National Natural Science Foundation of China
- 2014ZX09201042/National Mega-project for Innovative Drugs
- 2016-I2M-2-002/the CAMS Innovation Fund for Medical Sciences (CIFMS)
- 2018ZX09711001-007-002/Drug Innovation Major Project of China
LinkOut - more resources
Full Text Sources
Research Materials

