Exploring Pharmacological Functions of Alternatively Spliced Variants of the Mu Opioid Receptor Gene, Oprm1, via Gene-Targeted Animal Models
- PMID: 35328429
- PMCID: PMC8950057
- DOI: 10.3390/ijms23063010
Exploring Pharmacological Functions of Alternatively Spliced Variants of the Mu Opioid Receptor Gene, Oprm1, via Gene-Targeted Animal Models
Abstract
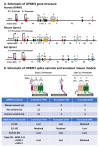
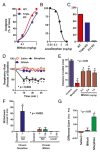
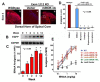
The mu opioid receptor has a distinct place in the opioid receptor family, since it mediates the actions of most opioids used clinically (e.g., morphine and fentanyl), as well as drugs of abuse (e.g., heroin). The single-copy mu opioid receptor gene, OPRM1, goes through extensive alternative pre-mRNA splicing to generate numerous splice variants that are conserved from rodents to humans. These OPRM1 splice variants can be classified into three structurally distinct types: (1) full-length 7 transmembrane (TM) carboxyl (C)-terminal variants; (2) truncated 6TM variants; and (3) single TM variants. Distinct pharmacological functions of these splice variants have been demonstrated by both in vitro and in vivo studies, particularly by using several unique gene-targeted mouse models. These studies provide new insights into our understanding of the complex actions of mu opioids with regard to OPRM1 alternative splicing. This review provides an overview of the studies that used these gene-targeted mouse models for exploring the functional importance of Oprm1 splice variants.
Keywords: OPRM1; alternative splicing; animal model; gene targeting; mu opioid receptor; opioid.
Conflict of interest statement
Y.-X.P. is a scientific co-founder of Sparian Biosciences. All other authors declare no conflict of interest.
Figures

References
-
- Martin W.R. Opioid Antagonists. Pharmacol. Rev. 1967;19:463–521. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

