Optimized Protocol for Proportionate CNS Cell Retrieval as a Versatile Platform for Cellular and Molecular Phenomapping in Aging and Neurodegeneration
- PMID: 35328432
- PMCID: PMC8950438
- DOI: 10.3390/ijms23063000
Optimized Protocol for Proportionate CNS Cell Retrieval as a Versatile Platform for Cellular and Molecular Phenomapping in Aging and Neurodegeneration
Abstract
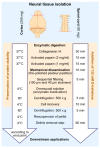
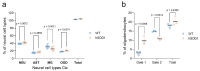
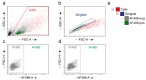
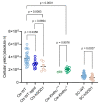
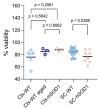
Efficient purification of viable neural cells from the mature CNS has been historically challenging due to the heterogeneity of the inherent cell populations. Moreover, changes in cellular interconnections, membrane lipid and cholesterol compositions, compartment-specific biophysical properties, and intercellular space constituents demand technical adjustments for cell isolation at different stages of maturation and aging. Though such obstacles are addressed and partially overcome for embryonic premature and mature CNS tissues, procedural adaptations to an aged, progeroid, and degenerative CNS environment are underrepresented. Here, we describe a practical workflow for the acquisition and phenomapping of CNS neural cells at states of health, physiological and precocious aging, and genetically provoked neurodegeneration. Following recent, unprecedented evidence of post-mitotic cellular senescence (PoMiCS), the protocol appears suitable for such de novo characterization and phenotypic opposition to classical senescence. Technically, the protocol is rapid, efficient as for cellular yield and well preserves physiological cell proportions. It is suitable for a variety of downstream applications aiming at cell type-specific interrogations, including cell culture systems, Flow-FISH, flow cytometry/FACS, senescence studies, and retrieval of omic-scale DNA, RNA, and protein profiles. We expect suitability for transfer to other CNS targets and to a broad spectrum of engineered systems addressing aging, neurodegeneration, progeria, and senescence.
Keywords: CNS aging; CNS cell isolation; amyotrophic lateral sclerosis (ALS); neurodegeneration; neuropathology; progeria; senescence.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
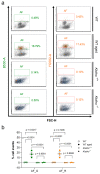

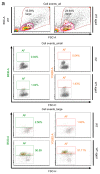

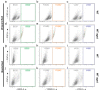
Similar articles
-
A genome-wide CRISPR-based screen identifies KAT7 as a driver of cellular senescence.Sci Transl Med. 2021 Jan 6;13(575):eabd2655. doi: 10.1126/scitranslmed.abd2655. Sci Transl Med. 2021. PMID: 33408182
-
Embryonic senescence and laminopathies in a progeroid zebrafish model.PLoS One. 2011 Mar 30;6(3):e17688. doi: 10.1371/journal.pone.0017688. PLoS One. 2011. PMID: 21479207 Free PMC article.
-
Reprogramming aging and progeria.Curr Opin Cell Biol. 2012 Dec;24(6):757-64. doi: 10.1016/j.ceb.2012.08.009. Epub 2012 Sep 6. Curr Opin Cell Biol. 2012. PMID: 22959961 Review.
-
Human aging and progeria.J Pediatr Endocrinol Metab. 2000;13 Suppl 6:1477-81. doi: 10.1515/jpem-2000-s622. J Pediatr Endocrinol Metab. 2000. PMID: 11202224 Review.
-
Erythrocyte Senescence in a Model of Rat Displaying Hutchinson-Gilford Progeria Syndrome.Anal Cell Pathol (Amst). 2018 May 29;2018:5028925. doi: 10.1155/2018/5028925. eCollection 2018. Anal Cell Pathol (Amst). 2018. PMID: 30003010 Free PMC article.
Cited by
-
Cellular Senescence in Physiological and Pathological Processes.Int J Mol Sci. 2022 Nov 1;23(21):13342. doi: 10.3390/ijms232113342. Int J Mol Sci. 2022. PMID: 36362123 Free PMC article.
References
MeSH terms
Grants and funding
- RegenerAging-52-5581-413-FSU-I-03/14 to A.K/Ministry for Economics, Sciences and Digital Society of Thuringia (TMWWDG), in frame of the ProExcellence Initiative RegenerAging
- PhD stipend to Q.A./Graduate Academy (GA) of the Jena University
- [FF01 to A.K.]/Interdisciplinary Centre for Clinical Research (IZKF) Jena
- [to O.W.W.]/Else-Kröner Research College 'Anti Age'
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

