Recent Advances in Structural Studies of Cytochrome bd and Its Potential Application as a Drug Target
- PMID: 35328590
- PMCID: PMC8951039
- DOI: 10.3390/ijms23063166
Recent Advances in Structural Studies of Cytochrome bd and Its Potential Application as a Drug Target
Abstract
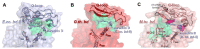
Cytochrome bd is a triheme copper-free terminal oxidase in membrane respiratory chains of prokaryotes. This unique molecular machine couples electron transfer from quinol to O2 with the generation of a proton motive force without proton pumping. Apart from energy conservation, the bd enzyme plays an additional key role in the microbial cell, being involved in the response to different environmental stressors. Cytochrome bd promotes virulence in a number of pathogenic species that makes it a suitable molecular drug target candidate. This review focuses on recent advances in understanding the structure of cytochrome bd and the development of its selective inhibitors.
Keywords: cytochrome oxidase; electron transport chain; enzyme structure; inhibition; membrane protein; molecular bioenergetics; terminal oxidase.
Conflict of interest statement
The authors declare that they have no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures





Similar articles
-
Oxygen as Acceptor.EcoSal Plus. 2015;6(2):10.1128/ecosalplus.ESP-0012-2015. doi: 10.1128/ecosalplus.ESP-0012-2015. EcoSal Plus. 2015. PMID: 26734697 Free PMC article. Review.
-
Cytochrome bd-type oxidases and environmental stressors in microbial physiology.Adv Microb Physiol. 2025;86:199-255. doi: 10.1016/bs.ampbs.2024.05.001. Epub 2024 Oct 30. Adv Microb Physiol. 2025. PMID: 40404270 Review.
-
Aerobic respiratory chain of Escherichia coli is not allowed to work in fully uncoupled mode.Proc Natl Acad Sci U S A. 2011 Oct 18;108(42):17320-4. doi: 10.1073/pnas.1108217108. Epub 2011 Oct 10. Proc Natl Acad Sci U S A. 2011. PMID: 21987791 Free PMC article.
-
[Cytochrome bd as Antioxidant Redox Enzyme].Mol Biol (Mosk). 2023 Nov-Dec;57(6):1084. Mol Biol (Mosk). 2023. PMID: 38062962 Review. Russian.
-
The terminal oxidase cytochrome bd-I confers carbon monoxide resistance to Escherichia coli cells.J Inorg Biochem. 2023 Oct;247:112341. doi: 10.1016/j.jinorgbio.2023.112341. Epub 2023 Jul 24. J Inorg Biochem. 2023. PMID: 37515940
Cited by
-
Identification of Small Molecule Inhibitors against Mycobacteria in Activated Macrophages.Molecules. 2022 Sep 8;27(18):5824. doi: 10.3390/molecules27185824. Molecules. 2022. PMID: 36144572 Free PMC article.
-
The cryoEM structure of cytochrome bd from C. glutamicum provides novel insights into structural properties of actinobacterial terminal oxidases.Front Chem. 2023 Jan 4;10:1085463. doi: 10.3389/fchem.2022.1085463. eCollection 2022. Front Chem. 2023. PMID: 36688035 Free PMC article.
-
Cytochrome bd oxidase: an emerging anti-tubercular drug target.RSC Med Chem. 2024 Jan 27;15(3):769-787. doi: 10.1039/d3md00587a. eCollection 2024 Mar 20. RSC Med Chem. 2024. PMID: 38516593 Free PMC article. Review.
-
Response of Mycobacterium smegmatis to the Cytochrome bcc Inhibitor Q203.Int J Mol Sci. 2022 Sep 7;23(18):10331. doi: 10.3390/ijms231810331. Int J Mol Sci. 2022. PMID: 36142240 Free PMC article.
-
Carbon Monoxide and Prokaryotic Energy Metabolism.Int J Mol Sci. 2025 Mar 20;26(6):2809. doi: 10.3390/ijms26062809. Int J Mol Sci. 2025. PMID: 40141451 Free PMC article. Review.
References
-
- Garcia-Bayona L., Coyne M.J., Hantman N., Montero-Llopis P., Von S.S., Ito T., Malamy M.H., Basler M., Barquera B., Comstock L.E. Nanaerobic growth enables direct visualization of dynamic cellular processes in human gut symbionts. Proc. Natl. Acad. Sci. USA. 2020;117:24484–24493. doi: 10.1073/pnas.2009556117. - DOI - PMC - PubMed
-
- Nikolaev A., Safarian S., Thesseling A., Wohlwend D., Friedrich T., Michel H., Kusumoto T., Sakamoto J., Melin F., Hellwig P. Electrocatalytic evidence of the diversity of the oxygen reaction in the bacterial bd oxidase from different organisms. Biochim. Biophys. Acta Bioenerg. 2021;1862:148436. doi: 10.1016/j.bbabio.2021.148436. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

