Comprehensive identification and expression analysis of CRY gene family in Gossypium
- PMID: 35331129
- PMCID: PMC8952943
- DOI: 10.1186/s12864-022-08440-9
Comprehensive identification and expression analysis of CRY gene family in Gossypium
Abstract
Background: The cryptochromes (CRY) are specific blue light receptors of plants and animals, which play crucial roles in physiological processes of plant growth, development, and stress tolerance.
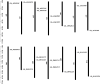
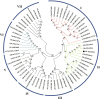
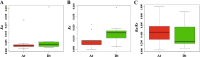
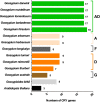
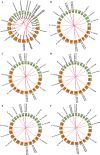
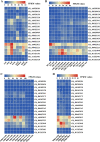
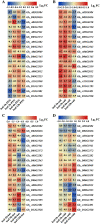
Results: In the present work, a systematic analysis of the CRY gene family was performed on twelve cotton species, resulting in 18, 17, 17, 17, and 17 CRYs identified in five alloteraploid cottons (Gossypium hirsutum, G. barbadense, G. tomentosum, G. mustelinum and G. darwinii), respectively, and five to nine CRY genes in the seven diploid species. Phylogenetic analysis of protein-coding sequences revealed that CRY genes from cottons and Arabidopsis thaliana could be classified into seven clades. Synteny analysis suggested that the homoeolog of G. hirsutum Gh_A02G0384 has undergone an evolutionary loss event in the other four allotetraploid cotton species. Cis-element analysis predicated the possible functions of CRY genes in G. hirsutum. RNA-seq data revealed that Gh_D09G2225, Gh_A09G2012 and Gh_A11G1040 had high expressions in fiber cells of different developmental states. In addition, the expression levels of one (Gh_A03G0120), 15 and nine GhCRY genes were down-regulated following the PEG, NaCl and high-temperature treatments, respectively. For the low-temperature treatment, five GhCRY genes were induced, and five were repressed. These results indicated that most GhCRY genes negatively regulate the abiotic stress treatments.
Conclusion: We report the structures, domains, divergence, synteny, and cis-elements analyses systematically of G. hirsutum CRY genes. Possible biological functions of GhCRY genes in differential tissues as well as in response to abiotic stress during the cotton plant life cycle were predicted.
Keywords: Abiotic stress; Cotton; Cryptochrome; Organ-specific expression; Phylogenetics.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

Similar articles
-
Comparative Genomics and Functional Studies of Putative m6A Methyltransferase (METTL) Genes in Cotton.Int J Mol Sci. 2022 Nov 15;23(22):14111. doi: 10.3390/ijms232214111. Int J Mol Sci. 2022. PMID: 36430588 Free PMC article.
-
Comprehensive characterization and gene expression patterns of LBD gene family in Gossypium.Planta. 2020 Mar 17;251(4):81. doi: 10.1007/s00425-020-03364-8. Planta. 2020. PMID: 32185507
-
Genome wide identification, classification and functional characterization of heat shock transcription factors in cultivated and ancestral cottons (Gossypium spp.).Int J Biol Macromol. 2021 Jul 1;182:1507-1527. doi: 10.1016/j.ijbiomac.2021.05.016. Epub 2021 May 7. Int J Biol Macromol. 2021. PMID: 33965497
-
Genome-wide identification of the MIOX gene family and their expression profile in cotton development and response to abiotic stress.PLoS One. 2021 Jul 9;16(7):e0254111. doi: 10.1371/journal.pone.0254111. eCollection 2021. PLoS One. 2021. PMID: 34242283 Free PMC article.
-
Genome-wide comparative analysis of NBS-encoding genes in four Gossypium species.BMC Genomics. 2017 Apr 12;18(1):292. doi: 10.1186/s12864-017-3682-x. BMC Genomics. 2017. PMID: 28403834 Free PMC article.
Cited by
-
Seed priming with graphene oxide improves salinity tolerance and increases productivity of peanut through modulating multiple physiological processes.J Nanobiotechnology. 2024 Sep 14;22(1):565. doi: 10.1186/s12951-024-02832-7. J Nanobiotechnology. 2024. PMID: 39272089 Free PMC article.
-
Comparative Genomics and Functional Studies of Putative m6A Methyltransferase (METTL) Genes in Cotton.Int J Mol Sci. 2022 Nov 15;23(22):14111. doi: 10.3390/ijms232214111. Int J Mol Sci. 2022. PMID: 36430588 Free PMC article.
-
Graphene oxide enhances aphid resistance in sorghum via the miR319-SbTCP7-SbLOX3 Pathway.Plant Biotechnol J. 2025 Aug;23(8):3164-3176. doi: 10.1111/pbi.70132. Epub 2025 May 21. Plant Biotechnol J. 2025. PMID: 40397692 Free PMC article.
-
AaMYC3 bridges the regulation of glandular trichome density and artemisinin biosynthesis in Artemisia annua.Plant Biotechnol J. 2025 Feb;23(2):315-332. doi: 10.1111/pbi.14449. Epub 2024 Aug 27. Plant Biotechnol J. 2025. PMID: 39189077 Free PMC article.
References
-
- Cashmore AR, Jarillo JA, Wu YJ, Liu D. Cryptochromes: blue light receptors for plants and animals. Science. 1999;284(5415):760–765. - PubMed
-
- Han X, Chang X, Zhang Z, Chen H, He H, Zhong B, Deng XW. Origin and evolution of core components responsible for monitoring light environment changes during plant terrestrialization. Mol Plant. 2019;12(6):847–862. - PubMed
-
- Ahmad M, Cashmore AR. HY4 gene of A. thaliana encodes a protein with characteristics of a blue-light photoreceptor. Nature. 1993;366(6451):162–166. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

