Re-evaluating the actin-dependence of spectraplakin functions during axon growth and maintenance
- PMID: 35333003
- PMCID: PMC9320987
- DOI: 10.1002/dneu.22873
Re-evaluating the actin-dependence of spectraplakin functions during axon growth and maintenance
Abstract
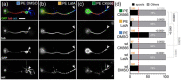
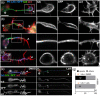
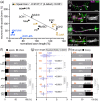
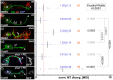
Axons are the long and slender processes of neurons constituting the biological cables that wire the nervous system. The growth and maintenance of axons require loose microtubule bundles that extend through their entire length. Understanding microtubule regulation is therefore an essential aspect of axon biology. Key regulators of neuronal microtubules are the spectraplakins, a well-conserved family of cytoskeletal cross-linkers that underlie neuropathies in mouse and humans. Spectraplakin deficiency in mouse or Drosophila causes severe decay of microtubule bundles and reduced axon growth. The underlying mechanisms are best understood for Drosophila's spectraplakin Short stop (Shot) and believed to involve cytoskeletal cross-linkage: Shot's binding to microtubules and Eb1 via its C-terminus has been thoroughly investigated, whereas its F-actin interaction via N-terminal calponin homology (CH) domains is little understood. Here, we have gained new understanding by showing that the F-actin interaction must be finely balanced: altering the properties of F-actin networks or deleting/exchanging Shot's CH domains induces changes in Shot function-with a Lifeact-containing Shot variant causing remarkable remodeling of neuronal microtubules. In addition to actin-microtubule (MT) cross-linkage, we find strong indications that Shot executes redundant MT bundle-promoting roles that are F-actin-independent. We argue that these likely involve the neuronal Shot-PH isoform, which is characterized by a large, unexplored central plakin repeat region (PRR) similarly existing also in mammalian spectraplakins.
Keywords: Drosophila; actin; axons; microtubules; neurons.
© 2022 The Authors. Developmental Neurobiology published by Wiley Periodicals LLC.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




References
-
- Alves‐Silva, J. , Sánchez‐Soriano, N. , Beaven, R. , Klein, M. , Parkin, J. , Millard, T. , Bellen, H. , Venken, K. J. T. , Ballestrem, C. , Kammerer, R. A. , & Prokop, A. (2012). Spectraplakins promote microtubule‐mediated axonal growth by functioning as structural microtubule‐associated proteins and EB1‐dependent +TIPs (Tip Interacting Proteins). Journal of Neuroscience, 32, 9143–9158. 10.1523/JNEUROSCI.0416-12.2012 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 087742/Z/08/Z/WT_/Wellcome Trust/United Kingdom
- BB/C515998/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- 077748/Z/05/Z/WT_/Wellcome Trust/United Kingdom
- 084561/Z/07/Z/WT_/Wellcome Trust/United Kingdom
- BB/R018960/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/M007553/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- 092403/Z/10/Z/WT_/Wellcome Trust/United Kingdom
- WT_/Wellcome Trust/United Kingdom
- BB/I002448/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- P40 OD018537/OD/NIH HHS/United States
- 087820/Z/08/Z/WT_/Wellcome Trust/United Kingdom
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

