Epidemiological and Genetic Characterization of Norovirus Outbreaks That Occurred in Catalonia, Spain, 2017-2019
- PMID: 35336893
- PMCID: PMC8955687
- DOI: 10.3390/v14030488
Epidemiological and Genetic Characterization of Norovirus Outbreaks That Occurred in Catalonia, Spain, 2017-2019
Abstract
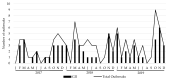
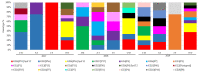
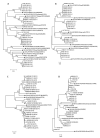
Molecular characterization of human norovirus (HuNoV) genotypes enhances the understanding of viral features and illustrates distinctive evolutionary patterns. The aim of our study was to describe the prevalence of the genetic diversity and the epidemiology of the genotypes involved in HuNoV outbreaks in Catalonia (Spain) between 2017 and 2019. A total of 100 HuNoV outbreaks were notified with the predominance of GII (70%), followed by GI (27%) and mixed GI/GII (3%). Seasonality was observed for GII outbreaks only. The most prevalent genotypes identified were GII.4[P31] Sydney 2012, GII.4[P16] Sydney 2012 and GII.2[P16]. As compared to person-to-person (P/P) transmitted outbreaks, foodborne outbreaks showed significantly higher attack rates and lower duration. The average attack rate was higher in youth hostel/campgrounds compared to nursing homes. Only genotypes GI.4[P4], GII.2[P16], GII.4[P16], GII.4[P31] and GII.17[P17] were consistently detected every year, and only abundance of GII.2[P16] showed a negative trend over time. GII.4 Sydney 2012 outbreaks were significantly associated to nursing homes, while GII.2[P16] and GI.3[P3] were most frequently identified in youth hostel/campgrounds. The average attack rate was significantly higher when comparing GII.2[P16] vs. GI.4[P4], GII.2[P16] vs. GII.4[P31] Sydney 2012, and GII.6[P7] vs. GII.4[P31] Sydney 2012. No correlations were found between genotype and outbreak duration or age of affected individuals.
Keywords: Spain; acute gastroenteritis outbreaks; genotyping; human norovirus; molecular epidemiology; viral diversity.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures

Similar articles
-
Molecular epidemiology and genetic diversity of norovirus infection in children hospitalized with acute gastroenteritis in East Java, Indonesia in 2015-2019.Infect Genet Evol. 2021 Mar;88:104703. doi: 10.1016/j.meegid.2020.104703. Epub 2021 Jan 2. Infect Genet Evol. 2021. PMID: 33401005
-
Recombinant Noroviruses Circulating in Spain from 2016 to 2020 and Proposal of Two Novel Genotypes within Genogroup I.Microbiol Spectr. 2022 Aug 31;10(4):e0250521. doi: 10.1128/spectrum.02505-21. Epub 2022 Jul 13. Microbiol Spectr. 2022. PMID: 35862999 Free PMC article.
-
Changing Predominance of Norovirus Recombinant Strains GII.2[P16] to GII.4[P16] and GII.4[P31] in Thailand, 2017 to 2018.Microbiol Spectr. 2022 Jun 29;10(3):e0044822. doi: 10.1128/spectrum.00448-22. Epub 2022 May 12. Microbiol Spectr. 2022. PMID: 35546545 Free PMC article.
-
Prevalence and Etiological Characteristics of Norovirus Infection in China: A Systematic Review and Meta-Analysis.Viruses. 2023 Jun 7;15(6):1336. doi: 10.3390/v15061336. Viruses. 2023. PMID: 37376635 Free PMC article.
-
Norovirus: An Overview of Virology and Preventative Measures.Viruses. 2022 Dec 16;14(12):2811. doi: 10.3390/v14122811. Viruses. 2022. PMID: 36560815 Free PMC article. Review.
Cited by
-
Epidemiological Characteristics of Norovirus Outbreaks in Shenyang from 2017 to 2021.J Microbiol. 2023 Apr;61(4):471-478. doi: 10.1007/s12275-023-00033-9. Epub 2023 Mar 27. J Microbiol. 2023. PMID: 36972002
-
Epidemiological Features of Human Norovirus Genotypes before and after COVID-19 Countermeasures in Osaka, Japan.Viruses. 2024 Apr 22;16(4):654. doi: 10.3390/v16040654. Viruses. 2024. PMID: 38675994 Free PMC article.
-
Biological and immunological characterization of major capsid protein VP1 from distinct GII.2 norovirus clusters.Sci Rep. 2024 Sep 9;14(1):21035. doi: 10.1038/s41598-024-72062-2. Sci Rep. 2024. PMID: 39251865 Free PMC article.
-
Molecular Evolution of RNA-Dependent RNA Polymerase Region in Norovirus Genogroup I.Viruses. 2023 Jan 5;15(1):166. doi: 10.3390/v15010166. Viruses. 2023. PMID: 36680206 Free PMC article.
-
Genetic Diversity and Phylogenetic Relationship of Human Norovirus Sequences Derived from Municipalities within the Sverdlovsk Region of Russia.Viruses. 2024 Jun 21;16(7):1001. doi: 10.3390/v16071001. Viruses. 2024. PMID: 39066164 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

