Bacterial Competition Systems Share a Domain Required for Inner Membrane Transport of the Bacteriocin Pyocin G from Pseudomonas aeruginosa
- PMID: 35343790
- PMCID: PMC9040868
- DOI: 10.1128/mbio.03396-21
Bacterial Competition Systems Share a Domain Required for Inner Membrane Transport of the Bacteriocin Pyocin G from Pseudomonas aeruginosa
Abstract
Bacteria exploit a variety of attack strategies to gain dominance within ecological niches. Prominent among these are contact-dependent inhibition (CDI), type VI secretion (T6SS), and bacteriocins. The cytotoxic endpoint of these systems is often the delivery of a nuclease to the cytosol. How such nucleases translocate across the cytoplasmic membrane of Gram-negative bacteria is unknown. Here, we identify a small, conserved, 15-kDa domain, which we refer to as the inner membrane translocation (IMT) domain, that is common to T6SS and bacteriocins and linked to nuclease effector domains. Through fluorescence microscopy assays using intact and spheroplasted cells, we demonstrate that the IMT domain of the Pseudomonas aeruginosa-specific bacteriocin pyocin G (PyoG) is required for import of the toxin nuclease domain to the cytoplasm. We also show that translocation of PyoG into the cytosol is dependent on inner membrane proteins FtsH, a AAA+ATPase/protease, and TonB1, the latter more typically associated with transport of bacteriocins across the outer membrane. Our study reveals that the IMT domain directs the cytotoxic nuclease of PyoG to cross the cytoplasmic membrane and, more broadly, has been adapted for the transport of other toxic nucleases delivered into Gram-negative bacteria by both contact-dependent and contact-independent means. IMPORTANCE Nuclease bacteriocins are potential antimicrobials for the treatment of antibiotic-resistant bacterial infections. While the mechanism of outer membrane translocation is beginning to be understood, the mechanism of inner membrane transport is not known. This study uses PyoG as a model nuclease bacteriocin and defines a conserved domain that is essential for inner membrane translocation and is widespread in other bacterial competition systems. Additionally, the presented data link two membrane proteins, FtsH and TonB1, with inner membrane translocation of PyoG. These findings point to the general importance of this domain to the cellular uptake mechanisms of nucleases delivered by otherwise diverse and distinct bacterial competition systems. The work is also of importance for the design of new protein antibiotics.
Keywords: P. aeruginosa; antibiotic; bacterial competition; cell envelope; protein import; pyocin.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

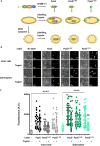


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
