Guide to development of compound files for PBPK modeling in the Simcyp population-based simulator
- PMID: 35344639
- PMCID: PMC9286711
- DOI: 10.1002/psp4.12791
Guide to development of compound files for PBPK modeling in the Simcyp population-based simulator
Abstract
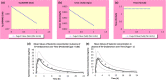
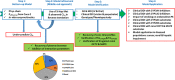
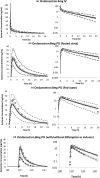
The Simcyp Simulator is a software platform for population physiologically-based pharmacokinetic (PBPK) modeling and simulation. It links in vitro data to in vivo absorption, distribution, metabolism, excretion and pharmacokinetic/pharmacodynamic outcomes to explore clinical scenarios and support drug development decisions, including regulatory submissions and drug labels. This tutorial describes the different input parameters required, as well as the considerations needed when developing a PBPK model within the Simulator, for a small molecule intended for oral administration. A case study showing the development and application of a PBPK model for ondansetron is herein used to aid the understanding of different PBPK model development concepts.
© 2022 The Authors. CPT: Pharmacometrics & Systems Pharmacology published by Wiley Periodicals LLC on behalf of American Society for Clinical Pharmacology and Therapeutics.
Conflict of interest statement
All authors are paid employees of Certara UK Limited (Simcyp Division) and may hold shares in Certara.
Figures

Similar articles
-
A guide to developing population files for physiologically-based pharmacokinetic modeling in the Simcyp Simulator.CPT Pharmacometrics Syst Pharmacol. 2024 Sep;13(9):1429-1447. doi: 10.1002/psp4.13202. Epub 2024 Jul 18. CPT Pharmacometrics Syst Pharmacol. 2024. PMID: 39030888 Free PMC article.
-
Using the Simcyp R Package for PBPK Simulation Workflows With the Simcyp Simulator.CPT Pharmacometrics Syst Pharmacol. 2025 May;14(5):853-863. doi: 10.1002/psp4.70022. Epub 2025 Apr 3. CPT Pharmacometrics Syst Pharmacol. 2025. PMID: 40179011 Free PMC article.
-
Does the choice of applied physiologically-based pharmacokinetics platform matter? A case study on simvastatin disposition and drug-drug interaction.CPT Pharmacometrics Syst Pharmacol. 2022 Sep;11(9):1194-1209. doi: 10.1002/psp4.12837. Epub 2022 Jul 16. CPT Pharmacometrics Syst Pharmacol. 2022. PMID: 35722750 Free PMC article.
-
Quality Assurance of PBPK Modeling Platforms and Guidance on Building, Evaluating, Verifying and Applying PBPK Models Prudently under the Umbrella of Qualification: Why, When, What, How and By Whom?Pharm Res. 2022 Aug;39(8):1733-1748. doi: 10.1007/s11095-022-03250-w. Epub 2022 Apr 20. Pharm Res. 2022. PMID: 35445350 Free PMC article. Review.
-
Predictive Pediatric Modeling and Simulation Using Ontogeny Information.J Clin Pharmacol. 2019 Sep;59 Suppl 1:S95-S103. doi: 10.1002/jcph.1497. J Clin Pharmacol. 2019. PMID: 31502689 Review.
Cited by
-
Key Factors for Improving Predictive Accuracy and Avoiding Overparameterization of the PBPK Absorption Model in Food Effect Studies of Weakly Basic Water-Insoluble Compounds in Immediate Release Formulations.Pharmaceutics. 2024 Oct 12;16(10):1324. doi: 10.3390/pharmaceutics16101324. Pharmaceutics. 2024. PMID: 39458653 Free PMC article.
-
PBPK modeling: What is the role of CYP3A4 expression in the gastrointestinal tract to accurately predict first-pass metabolism?CPT Pharmacometrics Syst Pharmacol. 2025 Jan;14(1):130-141. doi: 10.1002/psp4.13249. Epub 2024 Oct 2. CPT Pharmacometrics Syst Pharmacol. 2025. PMID: 39359052 Free PMC article.
-
Interspecies Brain PBPK Modeling Platform to Predict Passive Transport through the Blood-Brain Barrier and Assess Target Site Disposition.Pharmaceutics. 2024 Feb 4;16(2):226. doi: 10.3390/pharmaceutics16020226. Pharmaceutics. 2024. PMID: 38399280 Free PMC article.
-
Optimising Fluvoxamine Maternal/Fetal Exposure during Gestation: A Pharmacokinetic Virtual Clinical Trials Study.Metabolites. 2022 Dec 16;12(12):1281. doi: 10.3390/metabo12121281. Metabolites. 2022. PMID: 36557319 Free PMC article.
-
Development of a PBPK model to quantitatively understand absorption and disposition mechanism and support future clinical trials for PB-201.CPT Pharmacometrics Syst Pharmacol. 2023 Jul;12(7):941-952. doi: 10.1002/psp4.12964. Epub 2023 Apr 20. CPT Pharmacometrics Syst Pharmacol. 2023. PMID: 37078371 Free PMC article.
References
-
- Jamei M, Marciniak S, Feng K, Barnett A, Tucker G, Rostami‐Hodjegan A. The Simcyp population‐based ADME simulator. Expert Opin Drug Metab Toxicol. 2009;5:211‐223. - PubMed
-
- Zhao P, Rowland M, Huang SM. Best practice in the use of physiologically based pharmacokinetic modeling and simulation to address clinical pharmacology regulatory questions. Clin Pharmacol Ther. 2012;92:17‐20. - PubMed
-
- Beaumont K, Gardner I, Chapman K, Hall M, Rowland M. Toward an integrated human clearance prediction strategy that minimizes animal use. J Pharm Sci. 2011;100:4518‐4535. - PubMed
-
- Sandhya B, Ashwini HH, Ramesh KC, Seetharamappa J. Exploring the binding mechanism of ondansetron hydrochloride to serum albumins: spectroscopic approach. Spectrochim Acta A Mol Biomol Spectrosc. 2012;86:410‐416. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources

