Hybridization increases genetic diversity in Schistosoma haematobium populations infecting humans in Cameroon
- PMID: 35346375
- PMCID: PMC8962594
- DOI: 10.1186/s40249-022-00958-0
Hybridization increases genetic diversity in Schistosoma haematobium populations infecting humans in Cameroon
Abstract
Background: Hybrids between Schistosoma haematobium (Sh) and S. bovis (Sb) have been found in several African countries as well as in Europe. Since the consequences of this hybridization are still unknown, this study aims to verify the presence of such hybrids in Cameroonian humans, to describe the structure of S. haematobium populations on a large geographic scale, and to examine the impact of these hybrids on genetic diversity and structure of these populations.
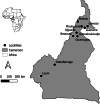
Methods: From January to April 2019, urine from infected children was collected in ten geographically distinct populations. Miracidia were collected from eggs in this urine. To detect the presence of hybrids among these miracidia we genotyped both Cox1 (RD-PCR) and ITS2 gene (PCR-RFLP). Population genetic diversity and structure was assessed by genotyping each miracidium with a panel of 14 microsatellite markers. Gene diversity was measured using both heterozygosity and allelic richness indexes, and genetic structure was analyzed using paired Fst, PCA and Bayesian approaches.
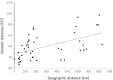
Results: Of the 1327 miracidia studied, 88.7% were identified as pure genotypes of S. haematobium (Sh_Sh/Sh) while the remaining 11.3% were hybrids (7.0% with Sh_Sh/Sb, 3.7% with Sb_Sb/Sh and 0.4% with Sb_Sh/Sb). No miracidium has been identified as a pure genotype of S. bovis. Allelic richness ranged from 5.55 (Loum population) to 7.73 (Matta-Barrage) and differed significantly between populations. Mean heterozygosity ranged from 53.7% (Loum) to 59% (Matta Barrage) with no significant difference. The overall genetic differentiation inferred either by a principal component analysis or by the Bayesian approach shows a partial structure. Southern populations (Loum and Matta Barrage) were clearly separated from other localities but genetic differentiation between northern localities was limited, certainly due to the geographic proximity between these sites.
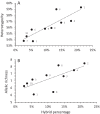
Conclusions: Hybrids between S. haematobium and S. bovis were identified in 11.3% of miracidia that hatched from eggs present in the urine of Cameroonian schoolchildren. The percentages of these hybrids are correlated with the genetic diversity of the parasite, indicating that hybridization increases genetic diversity in our sampling sites. Hybridization is therefore a major biological process that shapes the genetic diversity of S. haematobium.
Keywords: Cameroon; Genetic diversity; Hybridization; Miracidium; Schistosoma bovis; Schistosoma haematobium.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
References
-
- Harrison RG. Hybrid zones: windows on evolutionary process. In: Futuyma D, Antonovics J, editors. Oxford surveys in evolutionary biology. New York: Oxford University Press; 1990. pp. 69–128.
-
- Arnold ML. Natural hybridization as an evolutionary process. Annu Rev Ecol Syst. 1992;23:237–261. doi: 10.1146/annurev.es.23.110192.001321. - DOI
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

