Assessing the causal role of epigenetic clocks in the development of multiple cancers: a Mendelian randomization study
- PMID: 35346416
- PMCID: PMC9049976
- DOI: 10.7554/eLife.75374
Assessing the causal role of epigenetic clocks in the development of multiple cancers: a Mendelian randomization study
Abstract
Background: Epigenetic clocks have been associated with cancer risk in several observational studies. Nevertheless, it is unclear whether they play a causal role in cancer risk or if they act as a non-causal biomarker.
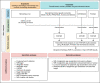
Methods: We conducted a two-sample Mendelian randomization (MR) study to examine the genetically predicted effects of epigenetic age acceleration as measured by HannumAge (nine single-nucleotide polymorphisms (SNPs)), Horvath Intrinsic Age (24 SNPs), PhenoAge (11 SNPs), and GrimAge (4 SNPs) on multiple cancers (i.e. breast, prostate, colorectal, ovarian and lung cancer). We obtained genome-wide association data for biological ageing from a meta-analysis (N = 34,710), and for cancer from the UK Biobank (N cases = 2671-13,879; N controls = 173,493-372,016), FinnGen (N cases = 719-8401; N controls = 74,685-174,006) and several international cancer genetic consortia (N cases = 11,348-122,977; N controls = 15,861-105,974). Main analyses were performed using multiplicative random effects inverse variance weighted (IVW) MR. Individual study estimates were pooled using fixed effect meta-analysis. Sensitivity analyses included MR-Egger, weighted median, weighted mode and Causal Analysis using Summary Effect Estimates (CAUSE) methods, which are robust to some of the assumptions of the IVW approach.
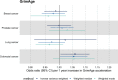
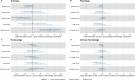
Results: Meta-analysed IVW MR findings suggested that higher GrimAge acceleration increased the risk of colorectal cancer (OR = 1.12 per year increase in GrimAge acceleration, 95% CI 1.04-1.20, p = 0.002). The direction of the genetically predicted effects was consistent across main and sensitivity MR analyses. Among subtypes, the genetically predicted effect of GrimAge acceleration was greater for colon cancer (IVW OR = 1.15, 95% CI 1.09-1.21, p = 0.006), than rectal cancer (IVW OR = 1.05, 95% CI 0.97-1.13, p = 0.24). Results were less consistent for associations between other epigenetic clocks and cancers.
Conclusions: GrimAge acceleration may increase the risk of colorectal cancer. Findings for other clocks and cancers were inconsistent. Further work is required to investigate the potential mechanisms underlying the results.
Funding: FMB was supported by a Wellcome Trust PhD studentship in Molecular, Genetic and Lifecourse Epidemiology (224982/Z/22/Z which is part of grant 218495/Z/19/Z). KKT was supported by a Cancer Research UK (C18281/A29019) programme grant (the Integrative Cancer Epidemiology Programme) and by the Hellenic Republic's Operational Programme 'Competitiveness, Entrepreneurship & Innovation' (OΠΣ 5047228). PH was supported by Cancer Research UK (C18281/A29019). RMM was supported by the NIHR Biomedical Research Centre at University Hospitals Bristol and Weston NHS Foundation Trust and the University of Bristol and by a Cancer Research UK (C18281/A29019) programme grant (the Integrative Cancer Epidemiology Programme). RMM is a National Institute for Health Research Senior Investigator (NIHR202411). The views expressed are those of the author(s) and not necessarily those of the NIHR or the Department of Health and Social Care. GDS and CLR were supported by the Medical Research Council (MC_UU_00011/1 and MC_UU_00011/5, respectively) and by a Cancer Research UK (C18281/A29019) programme grant (the Integrative Cancer Epidemiology Programme). REM was supported by an Alzheimer's Society project grant (AS-PG-19b-010) and NIH grant (U01 AG-18-018, PI: Steve Horvath). RCR is a de Pass Vice Chancellor's Research Fellow at the University of Bristol.
Keywords: DNA methylation; Mendelian randomization; cancer; epidemiology; epigenetic age acceleration; epigenetic clocks; genetics; genomics; human; medicine.
Plain language summary
Have you noticed that some people seem to get older faster than others? Scientists have previously found that a chemical tag on DNA known as DNA methylation can be used to predict an individual’s chronological age. However, age predicted using DNA methylation (also known as biological or epigenetic age) does not always perfectly correspond to chronological age. Indeed, some people’s biological age is higher than their years, while other people’s is lower. When an individual’s biological age is higher than their chronological age, they are said to be experiencing ‘epigenetic age acceleration’. This type of accelerated ageing, which can be measured with ‘epigenetic clocks’ based on DNA methylation, has been associated with several adverse health outcomes, including cancer. This means that epigenetic clocks may improve our ability to predict cancer risk and detect cancer early. However, it is still unclear whether accelerated biological ageing causes cancer, or whether it simply correlates with the disease. Morales-Berstein et al. wanted to investigate whether epigenetic age acceleration, as measured by epigenetic clocks, plays a role in the development of several cancers. To do so, they used an approach known as Mendelian randomization. Using genetic variants as natural experiments, they studied the effect of different measures of epigenetic age acceleration on cancer risk. Their work focused on five types of cancer: breast, colorectal, prostate, ovarian and lung cancer. They used genetic association data from people of European ancestry to determine whether genetic variants that are strongly associated with accelerated ageing are also strongly associated with cancer. The results showed that one of the DNA methylation markers used as an estimate of biological ageing could be directly related to the risk of developing colorectal cancer. This work provides new insights into the relationship between markers of biological ageing and cancer. Similar relationships should also be studied in other groups of people and for other cancer sites. The results suggest that reversing biological ageing by altering DNA methylation could prevent or delay the development of colorectal cancer.
© 2022, Morales Berstein et al.
Conflict of interest statement
FM, DM, KT, EB, PH, KB, AP, DB, IC, RM, GD, TR, RR No competing interests declared, AL declares that UC Regents filed the patent "DNA METHYLATION BASED BIOMARKERS FOR LIFE EXPECTANCY AND MORBIDITY" (International Application Number PCT/US2019/055444; in pending status) and that the Epigenetic Clock Development Foundation and Foxo Labs hold licenses, CR declares that UC Regents filed the patent "DNA METHYLATION BASED BIOMARKERS FOR LIFE EXPECTANCY AND MORBIDITY" (International Application Number PCT/US2019/055444; in pending status) and that the Epigenetic Clock Development Foundation and Foxo Labs hold licenses. SH receives consulting fees from the Epigenetic Clock Development Foundation and royalties for patents involving epigenetic clocks, SH has received a speaker fee from Illumina and is an advisor to the Epigenetic Clock Development Foundation, RM is employed part time by Novo Nordisk outside of this work
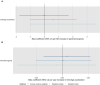
Figures




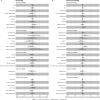


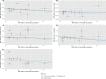

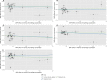






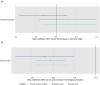



Comment in
- doi: 10.7554/eLife.78693
References
-
- Amos CI, Wu X, Broderick P, Gorlov IP, Gu J, Eisen T, Dong Q, Zhang Q, Gu X, Vijayakrishnan J, Sullivan K, Matakidou A, Wang Y, Mills G, Doheny K, Tsai Y-Y, Chen WV, Shete S, Spitz MR, Houlston RS. Genome-wide association scan of tag SNPs identifies a susceptibility locus for lung cancer at 15q25.1. Nature Genetics. 2008;40:616–622. doi: 10.1038/ng.109. - DOI - PMC - PubMed
-
- Benjamini Y, Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society. 1995;57:289–300. doi: 10.1111/j.2517-6161.1995.tb02031.x. - DOI
Publication types
MeSH terms
Grants and funding
- MC_QA137853/MRC_/Medical Research Council/United Kingdom
- C5047/A3354/CRUK_/Cancer Research UK/United Kingdom
- HHSN268201200008C/HL/NHLBI NIH HHS/United States
- R01 CA128978/CA/NCI NIH HHS/United States
- C1287/A 10710/CRUK_/Cancer Research UK/United Kingdom
- U19 CA148537/CA/NCI NIH HHS/United States
- C5047/A15007/CRUK_/Cancer Research UK/United Kingdom
- U19 CA148112/CA/NCI NIH HHS/United States
- C16913/A6135/CRUK_/Cancer Research UK/United Kingdom
- 29017/CRUK_/Cancer Research UK/United Kingdom
- U10 CA037429/CA/NCI NIH HHS/United States
- C5047/A7357/CRUK_/Cancer Research UK/United Kingdom
- C1287/A16563/CRUK_/Cancer Research UK/United Kingdom
- U01 CA188392/CA/NCI NIH HHS/United States
- C12292/A11174/CRUK_/Cancer Research UK/United Kingdom
- MC_UU_00011/1/MRC_/Medical Research Council/United Kingdom
- C5047/A10692/CRUK_/Cancer Research UK/United Kingdom
- 224982/Z/22/Z/WT_/Wellcome Trust/United Kingdom
- U19 CA148065/CA/NCI NIH HHS/United States
- WT_/Wellcome Trust/United Kingdom
- MC_PC_17228/MRC_/Medical Research Council/United Kingdom
- C1281/A12014/CRUK_/Cancer Research UK/United Kingdom
- C5047/A8384/CRUK_/Cancer Research UK/United Kingdom
- C8197/A16565/CRUK_/Cancer Research UK/United Kingdom
- HHSN268201200008I/HL/NHLBI NIH HHS/United States
- C1287/A10118/CRUK_/Cancer Research UK/United Kingdom
- UM1 CA182883/CA/NCI NIH HHS/United States
- MC_UU_00011/5/MRC_/Medical Research Council/United Kingdom
- C18281/A29019/CRUK_/Cancer Research UK/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

