Genetic variation of macronutrient tolerance in Drosophila melanogaster
- PMID: 35347148
- PMCID: PMC8960806
- DOI: 10.1038/s41467-022-29183-x
Genetic variation of macronutrient tolerance in Drosophila melanogaster
Abstract
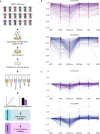
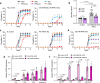
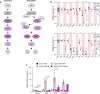
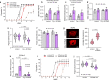
Carbohydrates, proteins and lipids are essential nutrients to all animals; however, closely related species, populations, and individuals can display dramatic variation in diet. Here we explore the variation in macronutrient tolerance in Drosophila melanogaster using the Drosophila genetic reference panel, a collection of ~200 strains derived from a single natural population. Our study demonstrates that D. melanogaster, often considered a "dietary generalist", displays marked genetic variation in survival on different diets, notably on high-sugar diet. Our genetic analysis and functional validation identify several regulators of macronutrient tolerance, including CG10960/GLUT8, Pkn and Eip75B. We also demonstrate a role for the JNK pathway in sugar tolerance and de novo lipogenesis. Finally, we report a role for tailless, a conserved orphan nuclear hormone receptor, in regulating sugar metabolism via insulin-like peptide secretion and sugar-responsive CCHamide-2 expression. Our study provides support for the use of nutrigenomics in the development of personalized nutrition.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
Mondo/ChREBP-Mlx-regulated transcriptional network is essential for dietary sugar tolerance in Drosophila.PLoS Genet. 2013 Apr;9(4):e1003438. doi: 10.1371/journal.pgen.1003438. Epub 2013 Apr 4. PLoS Genet. 2013. PMID: 23593032 Free PMC article.
-
Novel studies on Drosophila melanogaster model reveal the roles of JNK-Jak/STAT axis and intestinal microbiota in insulin resistance.J Drug Target. 2023 Mar;31(3):261-268. doi: 10.1080/1061186X.2022.2144869. Epub 2022 Nov 10. J Drug Target. 2023. PMID: 36343203 Review.
-
Physiological Adaptations to Sugar Intake: New Paradigms from Drosophila melanogaster.Trends Endocrinol Metab. 2017 Feb;28(2):131-142. doi: 10.1016/j.tem.2016.11.003. Epub 2016 Dec 5. Trends Endocrinol Metab. 2017. PMID: 27923532 Review.
-
Natural variation in sugar tolerance associates with changes in signaling and mitochondrial ribosome biogenesis.Elife. 2018 Nov 27;7:e40841. doi: 10.7554/eLife.40841. Elife. 2018. PMID: 30480548 Free PMC article.
-
A Single Nucleotide Variant in the PPARγ-homolog Eip75B Affects Fecundity in Drosophila.Mol Biol Evol. 2023 Feb 3;40(2):msad018. doi: 10.1093/molbev/msad018. Mol Biol Evol. 2023. PMID: 36703226 Free PMC article.
Cited by
-
Nutrigonometry IV: Thales' theorem to measure the rules of dietary compromise in animals.Sci Rep. 2023 May 8;13(1):7466. doi: 10.1038/s41598-023-34722-7. Sci Rep. 2023. PMID: 37156830 Free PMC article.
-
Drosophila as a Genetic Model System to Study Organismal Energy Metabolism.Biomolecules. 2025 May 1;15(5):652. doi: 10.3390/biom15050652. Biomolecules. 2025. PMID: 40427545 Free PMC article. Review.
-
Impact of Long-Term Pyriproxyfen Exposure on the Genetic Structure and Diversity of Aedes aegypti and Aedes albopictus in Manaus, Amazonas, Brazil.Genes (Basel). 2024 Aug 8;15(8):1046. doi: 10.3390/genes15081046. Genes (Basel). 2024. PMID: 39202406 Free PMC article.
-
Dhr96[1] mutation and maternal tudor[1] mutation increase life span and reduce the beneficial effects of mifepristone in mated female Drosophila.PLoS One. 2023 Dec 21;18(12):e0292820. doi: 10.1371/journal.pone.0292820. eCollection 2023. PLoS One. 2023. PMID: 38127988 Free PMC article.
-
Nutritional Trade-Offs in Drosophila melanogaster.Biology (Basel). 2025 Apr 7;14(4):384. doi: 10.3390/biology14040384. Biology (Basel). 2025. PMID: 40282249 Free PMC article.
References
-
- Havula E, Hietakangas V. Sugar sensing by ChREBP/Mondo-Mlx-new insight into downstream regulatory networks and integration of nutrient-derived signals. Curr. Opin. Cell Biol. 2018;51:89–96. - PubMed
-
- Mattila J, et al. Mondo-Mlx mediates organismal sugar sensing through the Gli-similar transcription factor sugarbabe. Cell Rep. 2015;13:350–364. - PubMed
-
- Havula, E. & Hietakangas, V. Glucose sensing by ChREBP/MondoA-Mlx transcription factors. Semin. Cell Dev. Biol.10.1016/j.semcdb.2012.02.007 (2012). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

