Secondary influenza challenge triggers resident memory B cell migration and rapid relocation to boost antibody secretion at infected sites
- PMID: 35349789
- PMCID: PMC9044924
- DOI: 10.1016/j.immuni.2022.03.003
Secondary influenza challenge triggers resident memory B cell migration and rapid relocation to boost antibody secretion at infected sites
Abstract
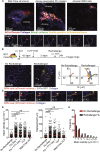
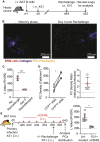
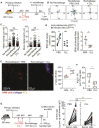
Resident memory B (BRM) cells develop and persist in the lungs of influenza-infected mice and humans; however, their contribution to recall responses has not been defined. Here, we used two-photon microscopy to visualize BRM cells within the lungs of influenza -virus immune and reinfected mice. Prior to re-exposure, BRM cells were sparsely scattered throughout the tissue, displaying limited motility. Within 24 h of rechallenge, these cells increased their migratory capacity, localized to infected sites, and subsequently differentiated into plasma cells. Alveolar macrophages mediated this process, in part by inducing expression of chemokines CXCL9 and CXCL10 from infiltrating inflammatory cells. This led to the recruitment of chemokine receptor CXCR3-expressing BRM cells to infected regions and increased local antibody concentrations. Our study uncovers spatiotemporal mechanisms that regulate lung BRM cell reactivation and demonstrates their capacity to rapidly deliver antibodies in a highly localized manner to sites of viral replication.
Keywords: antibody response; humoral immunity; influenza virus; live imaging; memory B cell; mucosal immunity; plasma cells; resident memory B cells; tissue-resident immunity; two-photon microscopy.
Copyright © 2022 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures





References
-
- Allen C.D., Okada T., Tang H.L., Cyster J.G. Imaging of germinal center selection events during affinity maturation. Science. 2007;315:528–531. - PubMed
-
- Alley M.R., Wells P.W., Smith W.D., Gardiner A.C. The distribution of immunoglobulin in the respiratory tract of sheep. Vet. Pathol. 1980;17:372–380. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

