PRMT7 ablation stimulates anti-tumor immunity and sensitizes melanoma to immune checkpoint blockade
- PMID: 35354055
- PMCID: PMC9838175
- DOI: 10.1016/j.celrep.2022.110582
PRMT7 ablation stimulates anti-tumor immunity and sensitizes melanoma to immune checkpoint blockade
Abstract
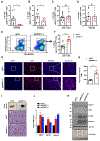
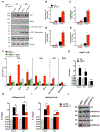
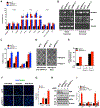
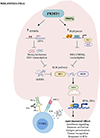
Despite the success of immune checkpoint inhibitor (ICI) therapy for cancer, resistance and relapse are frequent. Combination therapies are expected to enhance response rates and overcome this resistance. Herein, we report that combining PRMT7 inhibition with ICI therapy induces a strong anti-tumor T cell immunity and restrains tumor growth in vivo by increasing immune cell infiltration. PRMT7-deficient B16.F10 melanoma exhibits increased expression of genes in the interferon pathway, antigen presentation, and chemokine signaling. PRMT7 deficiency or inhibition with SGC3027 in B16.F10 melanoma results in reduced DNMT expression, loss of DNA methylation in the regulatory regions of endogenous retroviral elements (ERVs) causing their increased expression. PRMT7-deficient cells increase RIG-I and MDA5 expression with a reduction in the H4R3me2s repressive histone mark at their gene promoters. Our findings identify PRMT7 as a regulatory checkpoint for RIG-I, MDA5, and their ERV-double-stranded RNA (dsRNA) ligands, facilitating immune escape and anti-tumor T cell immunity to restrain tumor growth.
Keywords: CP: Cancer; CP: Immunology; DNMTs; PRMT7; RLR pathway; arginine methylation; cytotoxic T cells; dsRNA; immune checkpoint inhibitors; melanoma.
Copyright © 2022 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures




References
-
- Agolini E, Dentici ML, Bellacchio E, Alesi V, Radio FC, Torella A, Musacchia F, Tartaglia M, Dallapiccola B, Nigro V, et al. (2018). Expanding the clinical and molecular spectrum of PRMT7 mutations: 3 additional patients and review. Clin Genet 93, 675–681. - PubMed
-
- Balachandran S, Roberts PC, Brown LE, Truong H, Pattnaik AK, Archer DR, and Barber GN (2000). Essential role for the dsRNA-dependent protein kinase PKR in innate immunity to viral infection. Immunity 13, 129–141. - PubMed
-
- Benit L, De Parseval N, Casella JF, Callebaut I, Cordonnier A, and Heidmann T (1997). Cloning of a new murine endogenous retrovirus, MuERV-L, with strong similarity to the human HERV-L element and with a gag coding sequence closely related to the Fv1 restriction gene. Journal Of Virology 71, 5652–5657. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

