Biomimetic Lipopolysaccharide-Free Bacterial Outer Membrane-Functionalized Nanoparticles for Brain-Targeted Drug Delivery
- PMID: 35355446
- PMCID: PMC9165477
- DOI: 10.1002/advs.202105854
Biomimetic Lipopolysaccharide-Free Bacterial Outer Membrane-Functionalized Nanoparticles for Brain-Targeted Drug Delivery
Abstract
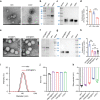
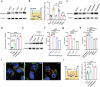
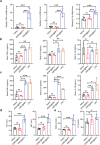
The blood-brain barrier (BBB) severely blocks the intracranial accumulation of most systemic drugs. Inspired by the contribution of the bacterial outer membrane to Escherichia coli K1 (EC-K1) binding to and invasion of BBB endothelial cells in bacterial meningitis, utilization of the BBB invasion ability of the EC-K1 outer membrane for brain-targeted drug delivery and construction of a biomimetic self-assembled nanoparticle with a surface featuring a lipopolysaccharide-free EC-K1 outer membrane are proposed. BBB penetration of biomimetic nanoparticles is demonstrated to occur through the transcellular vesicle transport pathway, which is at least partially dependent on internalization, endosomal escape, and transcytosis mediated by the interactions between outer membrane protein A and gp96 on BBB endothelial cells. This biomimetic nanoengineering strategy endows the loaded drugs with prolonged circulation, intracranial interstitial distribution, and extremely high biocompatibility. Based on the critical roles of gp96 in cancer biology, this strategy reveals enormous potential for delivering therapeutics to treat gp96-overexpressing intracranial malignancies.
Keywords: bacterial outer membrane; biomimetic; blood-brain barrier; drug delivery; nanoparticles.
© 2022 The Authors. Advanced Science published by Wiley-VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Escherichia coli interaction with human brain microvascular endothelial cells induces signal transducer and activator of transcription 3 association with the C-terminal domain of Ec-gp96, the outer membrane protein A receptor for invasion.Cell Microbiol. 2008 Nov;10(11):2326-38. doi: 10.1111/j.1462-5822.2008.01214.x. Epub 2008 Aug 15. Cell Microbiol. 2008. PMID: 18662321 Free PMC article.
-
A filter-free blood-brain barrier model to quantitatively study transendothelial delivery of nanoparticles by fluorescence spectroscopy.J Control Release. 2018 Nov 10;289:14-22. doi: 10.1016/j.jconrel.2018.09.015. Epub 2018 Sep 20. J Control Release. 2018. PMID: 30243824
-
DH5α Outer Membrane-Coated Biomimetic Nanocapsules Deliver Drugs to Brain Metastases but not Normal Brain Cells via Targeting GRP94.Small. 2023 Aug;19(35):e2300403. doi: 10.1002/smll.202300403. Epub 2023 Apr 27. Small. 2023. PMID: 37104822
-
Cell Membrane Biomimetic Nanoparticles with Potential in Treatment of Alzheimer's Disease.Molecules. 2023 Mar 3;28(5):2336. doi: 10.3390/molecules28052336. Molecules. 2023. PMID: 36903581 Free PMC article. Review.
-
Application of cell membrane-functionalized biomimetic nanoparticles in the treatment of glioma.J Mater Chem B. 2023 Aug 2;11(30):7055-7068. doi: 10.1039/d3tb00605k. J Mater Chem B. 2023. PMID: 37395053 Review.
Cited by
-
Opportunities and challenges of bacterial extracellular vesicles in regenerative medicine.J Nanobiotechnology. 2025 Jan 3;23(1):4. doi: 10.1186/s12951-024-02935-1. J Nanobiotechnology. 2025. PMID: 39754127 Free PMC article. Review.
-
Bacterial extracellular vesicles for gut microbiome-host communication and drug development.Acta Pharm Sin B. 2025 Apr;15(4):1816-1840. doi: 10.1016/j.apsb.2025.03.008. Epub 2025 Mar 7. Acta Pharm Sin B. 2025. PMID: 40486854 Free PMC article. Review.
-
Brain Delivery of IGF1R5, a Single-Domain Antibody Targeting Insulin-like Growth Factor-1 Receptor.Pharmaceutics. 2022 Jul 12;14(7):1452. doi: 10.3390/pharmaceutics14071452. Pharmaceutics. 2022. PMID: 35890347 Free PMC article.
-
Latest developments in biomaterial interfaces and drug delivery: challenges, innovations, and future outlook.Z Naturforsch C J Biosci. 2024 Nov 21. doi: 10.1515/znc-2024-0208. Online ahead of print. Z Naturforsch C J Biosci. 2024. PMID: 39566511 Review.
-
Gene editing tool-loaded biomimetic cationic vesicles with highly efficient bacterial internalization for in vivo eradication of pathogens.J Nanobiotechnology. 2024 Dec 22;22(1):787. doi: 10.1186/s12951-024-03065-4. J Nanobiotechnology. 2024. PMID: 39710679 Free PMC article.
References
-
- a) Ben‐Zvi A., Lacoste B., Kur E., Andreone B. J., Mayshar Y., Yan H., Gu C., Nature 2014, 509, 507; - PMC - PubMed
- b) Andreone B. J., Chow B. W., Tata A., Lacoste B., Ben‐Zvi A., Bullock K., Deik A. A., Ginty D. D., Clish C. B., Gu C., Neuron 2017, 94, 581; - PMC - PubMed
- c) Wang J. Z., Xiao N., Zhang Y. Z., Zhao C. X., Guo X. H., Lu L. M., Pharmacol Res 2016, 104, 124. - PubMed
-
- a) Anraku Y., Kuwahara H., Fukusato Y., Mizoguchi A., Ishii T., Nitta K., Matsumoto Y., Toh K., Miyata K., Uchida S., Nishina K., Osada K., Itaka K., Nishiyama N., Mizusawa H., Yamasoba T., Yokota T., Kataoka K., Nat. Commun. 2017, 8, 1001; - PMC - PubMed
- b) Niu X., Chen J., Gao J., Asian J Pharm. Sci. 2019, 14, 480; - PMC - PubMed
- c) Wang H., Zhang Z., Guan J., Lu W., Zhan C., Asian J Pharm. Sci. 2021, 16, 120. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 81973254/National Natural Science Foundation of China
- 32171381/National Natural Science Foundation of China
- BK20191421/Natural Science Foundation of Jiangsu Province
- SYS2019033/Suzhou Science and Technology Development Project
- Priority Academic Program Development of the Jiangsu Higher Education Institutes
LinkOut - more resources
Full Text Sources
