Profile and Functional Prediction of Plasma Exosome-Derived CircRNAs From Acute Ischemic Stroke Patients
- PMID: 35360855
- PMCID: PMC8963851
- DOI: 10.3389/fgene.2022.810974
Profile and Functional Prediction of Plasma Exosome-Derived CircRNAs From Acute Ischemic Stroke Patients
Abstract
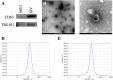
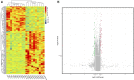
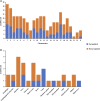
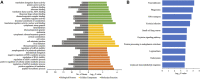
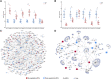
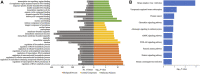
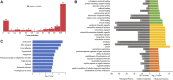
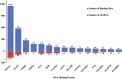
Stroke is one of the major causes of death and long-term disability, of which acute ischemic stroke (AIS) is the most common type. Although circular RNA (circRNA) expression profiles of AIS patients have been reported to be significantly altered in blood and peripheral blood mononuclear cells, the role of exosome-containing circRNAs after AIS is still unknown. Plasma exosomes from 10 AIS patients and 10 controls were isolated, and through microarray and bioinformatics analysis, the profile and putative function of circRNAs in the plasma exosomes were studied. A total of 198 circRNAs were differentially quantified (|log2 fold change| ≥ 1.00, p < 0.05) between AIS patients and controls. The levels of 12 candidate circRNAs were verified by qRT-PCR, and the quantities of 10 of these circRNAs were consistent with the data of microarray. The functions of host genes of differentially quantified circRNAs, including RNA and protein process, focal adhesion, and leukocyte transendothelial migration, were associated with the development of AIS. As a miRNA sponge, differentially quantified circRNAs had the potential to regulate pathways related to AIS, like PI3K-Akt, AMPK, and chemokine pathways. Of 198 differentially quantified circRNAs, 96 circRNAs possessing a strong translational ability could affect cellular structure and activity, like focal adhesion, tight junction, and endocytosis. Most differentially quantified circRNAs were predicted to bind to EIF4A3 and AGO2-two RNA-binding proteins (RBPs)-and to play a role in AIS. Moreover, four of ten circRNAs with verified levels by qRT-PCR (hsa_circ_0112036, hsa_circ_0066867, hsa_circ_0093708, and hsa_circ_0041685) were predicted to participate in processes of AIS, including PI3K-Akt, AMPK, and chemokine pathways as well as endocytosis, and to be potentially useful as diagnostic biomarkers for AIS. In conclusion, plasma exosome-derived circRNAs were significantly differentially quantified between AIS patients and controls and participated in the occurrence and progression of AIS by sponging miRNA/RBPs or translating into proteins, indicating that circRNAs from plasma exosomes could be crucial molecules in the pathogenesis of AIS and promising candidates as diagnostic biomarkers and therapeutic targets for the condition.
Keywords: acute ischemic stroke; ceRNA; circRNA; plasma exosome; profile.
Copyright © 2022 Yang, Hao, Lin, Guo, Liao, Yang, Cheng, Yang and Chen.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

