Altered propionate metabolism contributes to tumour progression and aggressiveness
- PMID: 35361954
- PMCID: PMC9050834
- DOI: 10.1038/s42255-022-00553-5
Altered propionate metabolism contributes to tumour progression and aggressiveness
Abstract
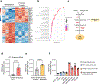
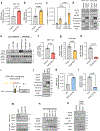
The alteration of metabolic pathways is a critical strategy for cancer cells to attain the traits necessary for metastasis in disease progression. Here, we find that dysregulation of propionate metabolism produces a pro-aggressive signature in breast and lung cancer cells, increasing their metastatic potential. This occurs through the downregulation of methylmalonyl coenzyme A epimerase (MCEE), mediated by an extracellular signal-regulated kinase 2-driven transcription factor Sp1/early growth response protein 1 transcriptional switch driven by metastatic signalling at its promoter level. The loss of MCEE results in reduced propionate-driven anaplerotic flux and intracellular and intratumoral accumulation of methylmalonic acid, a by-product of propionate metabolism that promotes cancer cell invasiveness. Altogether, we present a previously uncharacterized dysregulation of propionate metabolism as an important contributor to cancer and a valuable potential target in the therapeutic treatment of metastatic carcinomas.
© 2022. The Author(s), under exclusive licence to Springer Nature Limited.
Figures

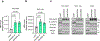






Comment in
-
Methylmalonic acid: an age-related metabolite that drives tumour aggressiveness.Nat Metab. 2022 Apr;4(4):412-413. doi: 10.1038/s42255-022-00540-w. Nat Metab. 2022. PMID: 35361957 No abstract available.
References
-
- Mortality GBD & Causes of Death C Global, regional, and national life expectancy, all-cause mortality, and cause-specific mortality for 249 causes of death, 1980–2015: a systematic analysis for the Global Burden of Disease Study 2015. Lancet 388, 1459–1544, doi:10.1016/S0140-6736(16)31012-1 (2016). - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

