Green Banana Flour Contributes to Gut Microbiota Recovery and Improves Colonic Barrier Integrity in Mice Following Antibiotic Perturbation
- PMID: 35369097
- PMCID: PMC8964434
- DOI: 10.3389/fnut.2022.832848
Green Banana Flour Contributes to Gut Microbiota Recovery and Improves Colonic Barrier Integrity in Mice Following Antibiotic Perturbation
Abstract
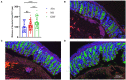
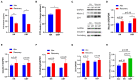
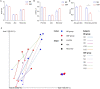
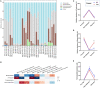
Green banana flour (GBF) is rich in resistant starch that has been used as a prebiotic to exert beneficial effects on gut microbiota. In this study, GBF was evaluated for its capacity to restore gut microbiota and intestinal barrier integrity from antibiotics (Abx) perturbation by comparing it to natural recovery (NR) treatment. C57B/L 6 J mice were exposed to 3 mg ciprofloxacin and 3.5 mg metronidazole once a day for 2 weeks to induce gut microbiota dysbiosis model. Then, GBF intervention at the dose of 400 mg/kg body weight was conducted for 2 weeks. The results showed that mice treated with Abx displayed increased gut permeability and intestinal barrier disruption, which were restored more quickly with GBF than NR treatment by increasing the secretion of mucin. Moreover, GBF treatment enriched beneficial Bacteroidales S24-7, Lachnospiraceae, Bacteroidaceae, and Porphyromonadaceae that accelerated the imbalanced gut microbiota restoration to its original state. This study puts forward novel insights into the application of GBF as a functional food ingredient to repair gut microbiota from Abx perturbation.
Keywords: antibiotic; green banana flour; gut microbiota; intestinal barrier; recovery.
Copyright © 2022 Li, Li, Song, Huang, Wu, Xu and Lu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

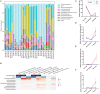

References
LinkOut - more resources
Full Text Sources

