Development of a sensitive trial-ready poly(GP) CSF biomarker assay for C9orf72-associated frontotemporal dementia and amyotrophic lateral sclerosis
- PMID: 35379698
- PMCID: PMC9279742
- DOI: 10.1136/jnnp-2021-328710
Development of a sensitive trial-ready poly(GP) CSF biomarker assay for C9orf72-associated frontotemporal dementia and amyotrophic lateral sclerosis
Abstract
Objective: A GGGGCC repeat expansion in the C9orf72 gene is the most common cause of genetic frontotemporal dementia (FTD) and amyotrophic lateral sclerosis (ALS). As potential therapies targeting the repeat expansion are now entering clinical trials, sensitive biomarker assays of target engagement are urgently required. Our objective was to develop such an assay.
Methods: We used the single molecule array (Simoa) platform to develop an immunoassay for measuring poly(GP) dipeptide repeat proteins (DPRs) generated by the C9orf72 repeat expansion in cerebrospinal fluid (CSF) of people with C9orf72-associated FTD/ALS.
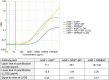
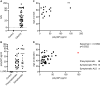
Results and conclusions: We show the assay to be highly sensitive and robust, passing extensive qualification criteria including low intraplate and interplate variability, a high precision and accuracy in measuring both calibrators and samples, dilutional parallelism, tolerance to sample and standard freeze-thaw and no haemoglobin interference. We used this assay to measure poly(GP) in CSF samples collected through the Genetic FTD Initiative (N=40 C9orf72 and 15 controls). We found it had 100% specificity and 100% sensitivity and a large window for detecting target engagement, as the C9orf72 CSF sample with the lowest poly(GP) signal had eightfold higher signal than controls and on average values from C9orf72 samples were 38-fold higher than controls, which all fell below the lower limit of quantification of the assay. These data indicate that a Simoa-based poly(GP) DPR assay is suitable for use in clinical trials to determine target engagement of therapeutics aimed at reducing C9orf72 repeat-containing transcripts.
Keywords: FRONTOTEMPORAL DEMENTIA; MOTOR NEURON DISEASE.
© Author(s) (or their employer(s)) 2022. Re-use permitted under CC BY. Published by BMJ.
Conflict of interest statement
Competing interests: SP, SM, YL, JG and RB were paid employees of Wave Life Sciences during completion of this work. JDR is on a Medical Advisory Board for Wave Life Sciences. JMS has received research funding from Avid Radiopharmaceuticals (a wholly owned subsidiary of Eli Lilly), has consulted for Roche Pharmaceuticals, Biogen, Merck and Eli Lilly, given educational lectures sponsored by GE Healthcare, Eli Lilly, and Biogen, and serves on a Data Safety Monitoring Committee for Axon Neuroscience SE. HZ has served at scientific advisory boards for Alector, Eisai, Denali, Roche Diagnostics, Wave, Samumed, Siemens Healthineers, Pinteon Therapeutics, Nervgen, AZTherapies and CogRx, has given lectures in symposia sponsored by Cellectricon, Fujirebio, Alzecure and Biogen, and is a co-founder of Brain Biomarker Solutions in Gothenburg AB (BBS), which is a part of the GU Ventures Incubator Program (outside submitted work). JBR has provided consultancy unrelated to the current work for Asceneuron, Astex, Biogen, UCB, SV Health, Curasen.
Figures


References
-
- van Blitterswijk M, DeJesus-Hernandez M, Niemantsverdriet E, et al. . Association between repeat sizes and clinical and pathological characteristics in carriers of C9orf72 repeat expansions (Xpansize-72): a cross-sectional cohort study. Lancet Neurol 2013;12:978–88. 10.1016/S1474-4422(13)70210-2 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- MC_UU_00030/14/MRC_/Medical Research Council/United Kingdom
- MC_U105597119/MRC_/Medical Research Council/United Kingdom
- MALASPINA/APR13/817-791/MNDA_/Motor Neurone Disease Association/United Kingdom
- MR/M023664/1/MRC_/Medical Research Council/United Kingdom
- WT_/Wellcome Trust/United Kingdom
- MC_UU_00005/12/MRC_/Medical Research Council/United Kingdom
- TURNER/OCT15/972-797/MNDA_/Motor Neurone Disease Association/United Kingdom
- MR/M008525/1/MRC_/Medical Research Council/United Kingdom
- PG/17/90/33415/BHF_/British Heart Foundation/United Kingdom
- MR/T046015/1/MRC_/Medical Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous
