Regional Salmonella Differences in United States Broiler Production from 2016 to 2020 and the Contribution of Multiserovar Populations to Salmonella Surveillance
- PMID: 35384708
- PMCID: PMC9040615
- DOI: 10.1128/aem.00204-22
Regional Salmonella Differences in United States Broiler Production from 2016 to 2020 and the Contribution of Multiserovar Populations to Salmonella Surveillance
Abstract
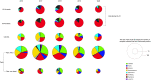
Poultry remains a considerable source of foodborne salmonellosis despite significant reduction of Salmonella incidence during processing. There are multiple entry points for Salmonella during production that can lead to contamination during slaughter, and it is important to distinguish the serovars present between the different stages to enact appropriate controls. National Salmonella data from the U.S. Department of Agriculture-Food Safety Inspection Service (USDA-FSIS) monitoring of poultry processing was analyzed from 2016 to 2020. The overall Salmonella incidence at processing in broiler carcasses and intact parts (parts) decreased from 9.00 to 6.57% over this period. The incidence in parts was higher (11.15%) than in carcasses (4.78%). Regional differences include higher proportions of serovars Infantis and Typhimurium in the Atlantic and higher proportion of serovar Schwarzengrund in the Southeast. For Georgia, the largest broiler-producing state, USDA-FSIS data were compared to Salmonella monitoring data from breeder flocks over the same period, revealing serovar Kentucky as the major serovar in breeders (67.91%) during production but not at processing, suggesting that it is more effectively removed during antimicrobial interventions. CRISPR-SeroSeq was performed on breeder samples collected between 2020 and 2021 to explain the incongruence between pre- and postharvest and showed that 32% of samples contain multiple serovars, with up to 11 serovars found in a single flock. High-resolution sequencing identifies serovar patterns at the population level and can provide insight to develop targeted controls. The work presented may apply to other food production systems where Salmonella is a concern, since it overcomes limitations associated with conventional culture. IMPORTANCE Salmonella is a leading cause of bacterial foodborne illness in the United States, with poultry as a significant Salmonella reservoir. We show the relative decrease in Salmonella over a 5-year period from 2016 to 2020 in processed chicken parts and highlight regional differences with respect to the prevalence of clinically important Salmonella serovars. Our results show that the discrepancy between Salmonella serovars found in pre- and postharvest poultry during surveillance are due in part by the limited detection depth offered by traditional culture techniques. Despite the reduction of Salmonella at processing, the number of human salmonellosis cases has remained stable, which may be attributed to differences in virulence among serovars and their associated risk. When monitoring for Salmonella, it is imperative to identify all serovars present to appropriately assess public health risk and to implement the most effective Salmonella controls.
Keywords: CRISPR-SeroSeq; Salmonella; monitoring; poultry; salmonellosis; serovars.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Combined Quantification and Deep Serotyping for Salmonella Risk Profiling in Broiler Flocks.Appl Environ Microbiol. 2023 Apr 26;89(4):e0203522. doi: 10.1128/aem.02035-22. Epub 2023 Mar 15. Appl Environ Microbiol. 2023. PMID: 36920215 Free PMC article.
-
High-Resolution Identification of Multiple Salmonella Serovars in a Single Sample by Using CRISPR-SeroSeq.Appl Environ Microbiol. 2018 Oct 17;84(21):e01859-18. doi: 10.1128/AEM.01859-18. Print 2018 Nov 1. Appl Environ Microbiol. 2018. PMID: 30170999 Free PMC article.
-
Dominant Salmonella Serovars in Australian Broiler Breeder Flocks and Hatcheries: a Longitudinal Study.Appl Environ Microbiol. 2023 Aug 30;89(8):e0062723. doi: 10.1128/aem.00627-23. Epub 2023 Jul 19. Appl Environ Microbiol. 2023. PMID: 37466445 Free PMC article.
-
Salmonella challenges: prevalence in swine and poultry and potential pathogenicity of such isolates.J Anim Sci. 2008 Apr;86(14 Suppl):E149-62. doi: 10.2527/jas.2007-0464. Epub 2007 Oct 2. J Anim Sci. 2008. PMID: 17911227 Review.
-
Impact of sporadic reporting of poultry Salmonella serovars from selected developing countries.J Infect Dev Ctries. 2015 Jan 15;9(1):1-7. doi: 10.3855/jidc.5065. J Infect Dev Ctries. 2015. PMID: 25596565 Review.
Cited by
-
Prevalence and molecular characterization of Salmonella isolated from wild birds in fresh produce environments.Front Microbiol. 2023 Nov 7;14:1272916. doi: 10.3389/fmicb.2023.1272916. eCollection 2023. Front Microbiol. 2023. PMID: 38029194 Free PMC article.
-
Genomic and phenotypic characterization of Salmonella enterica serovar Kentucky.Microb Genom. 2023 Sep;9(9):001089. doi: 10.1099/mgen.0.001089. Microb Genom. 2023. PMID: 37750759 Free PMC article.
-
Salmonella Biofilm Formation under Fluidic Shear Stress on Different Surface Materials.Foods. 2023 May 8;12(9):1918. doi: 10.3390/foods12091918. Foods. 2023. PMID: 37174455 Free PMC article.
-
Emergence and characteristics of multidrug-resistant Salmonella enterica subspecies enterica serovar Infantis harboring the pESI plasmid in chicken slaughterhouses in South Korea.Microbiol Spectr. 2025 Jul;13(7):e0295524. doi: 10.1128/spectrum.02955-24. Epub 2025 Jun 2. Microbiol Spectr. 2025. PMID: 40454925 Free PMC article.
-
Genetic Characteristics of Salmonella Isolates Recovered From Reused Broiler Litter Over Three Successive Flocks.J Food Prot. 2024 Mar;87(3):100236. doi: 10.1016/j.jfp.2024.100236. Epub 2024 Feb 1. J Food Prot. 2024. PMID: 38307462 Free PMC article.
References
-
- Interagency Food Safety Analytics Collaboration. 2021. Foodborne illness source attribution estimates for 2019 for Salmonella, Escherichia coli O157, Listeria monocytogenes, and Campylobacter using multi-year outbreak surveillance data, United States. https://www.fda.gov/food/cfsan-constituent-updates/release-2019-annual-r.... Accessed 15 January 2022.
-
- Grimont P, Weill F-X. 2007. Antigenic formulae of the Salmonella serovars: WHO Collaborating Centre for Reference and Research on Salmonella, p 1–166, 9th ed. Institute Pasteur, Paris, France.
-
- Cohn AR, Cheng RA, Orsi RH, Wiedmann M. 2021. Moving past species classifications for risk-based approaches to food safety: Salmonella as a case study. Front Sustain Food Syst 5:652132. 10.3389/fsufs.2021.652132. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

