Direct measurements of mRNA translation kinetics in living cells
- PMID: 35388013
- PMCID: PMC8986856
- DOI: 10.1038/s41467-022-29515-x
Direct measurements of mRNA translation kinetics in living cells
Abstract
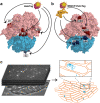
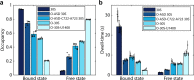
Ribosome mediated mRNA translation is central to life. The cycle of translation, however, has been characterized mostly using reconstituted systems, with only few techniques applicable for studies in the living cell. Here we describe a live-cell ribosome-labeling method, which allows us to characterize the whole processes of finding and translating an mRNA, using single-molecule tracking techniques. We find that more than 90% of both bacterial ribosomal subunits are engaged in translation at any particular time, and that the 30S and 50S ribosomal subunits spend the same average time bound to an mRNA, revealing that 30S re-initiation on poly-cistronic mRNAs is not prevalent in E. coli. Instead, our results are best explained by substantial 70S re-initiation of translation of poly-cistronic mRNAs, which is further corroborated by experiments with translation initiation inhibitors. Finally, we find that a variety of previously described orthogonal ribosomes, with altered anti-Shine-Dalgarno sequences, show significant binding to endogenous mRNAs.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





Similar articles
-
Translation of mRNAs with degenerate initiation triplet AUU displays high initiation factor 2 dependence and is subject to initiation factor 3 repression.Proc Natl Acad Sci U S A. 1993 May 1;90(9):4161-5. doi: 10.1073/pnas.90.9.4161. Proc Natl Acad Sci U S A. 1993. PMID: 8483930 Free PMC article.
-
Translation enhancer improves the ribosome liberation from translation initiation.J Am Chem Soc. 2013 Sep 4;135(35):13096-106. doi: 10.1021/ja405967h. Epub 2013 Aug 22. J Am Chem Soc. 2013. PMID: 23927491
-
Leaderless mRNAs bind 70S ribosomes more strongly than 30S ribosomal subunits in Escherichia coli.J Bacteriol. 2002 Dec;184(23):6730-3. doi: 10.1128/JB.184.23.6730-6733.2002. J Bacteriol. 2002. PMID: 12426363 Free PMC article.
-
The diversity of Shine-Dalgarno sequences sheds light on the evolution of translation initiation.RNA Biol. 2021 Nov;18(11):1489-1500. doi: 10.1080/15476286.2020.1861406. Epub 2020 Dec 21. RNA Biol. 2021. PMID: 33349119 Free PMC article. Review.
-
Specialized ribosomes in Escherichia coli.Biotechnol Prog. 1993 Sep-Oct;9(5):443-9. doi: 10.1021/bp00023a001. Biotechnol Prog. 1993. PMID: 7764160 Review.
Cited by
-
Spatiotemporal kinetics of the SRP pathway in live E. coli cells.Proc Natl Acad Sci U S A. 2022 Sep 20;119(38):e2204038119. doi: 10.1073/pnas.2204038119. Epub 2022 Sep 12. Proc Natl Acad Sci U S A. 2022. PMID: 36095178 Free PMC article.
-
Intradermal Delivery of Naked mRNA Vaccines via Iontophoresis.Pharmaceutics. 2023 Nov 26;15(12):2678. doi: 10.3390/pharmaceutics15122678. Pharmaceutics. 2023. PMID: 38140019 Free PMC article. Review.
-
ExTrack characterizes transition kinetics and diffusion in noisy single-particle tracks.J Cell Biol. 2023 May 1;222(5):e202208059. doi: 10.1083/jcb.202208059. Epub 2023 Feb 28. J Cell Biol. 2023. PMID: 36880553 Free PMC article.
-
Dynamic binding of the bacterial chaperone Trigger factor to translating ribosomes in Escherichia coli.Proc Natl Acad Sci U S A. 2025 Jan 7;122(1):e2409536121. doi: 10.1073/pnas.2409536121. Epub 2024 Dec 31. Proc Natl Acad Sci U S A. 2025. PMID: 39739798 Free PMC article.
-
A complex between IF2 and NusA suggests early coupling of transcription-translation.Nat Commun. 2025 Jul 26;16(1):6906. doi: 10.1038/s41467-025-62207-w. Nat Commun. 2025. PMID: 40715203 Free PMC article.
References
-
- Elf J, Barkefors I. Single-molecule kinetics in living cells. Annu. Rev. Biochem. 2019;88:635–659. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

