An Evolutionary Insight Into the Heterogeneous Severity Pattern of the SARS-CoV-2 Infection
- PMID: 35391792
- PMCID: PMC8981084
- DOI: 10.3389/fgene.2022.859508
An Evolutionary Insight Into the Heterogeneous Severity Pattern of the SARS-CoV-2 Infection
Abstract
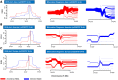
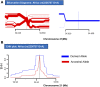
The ongoing pandemic of COVID-19 has elaborated an idiosyncratic pattern of SARS-CoV-2-induced symptoms in the human host. Some populations have succumbed to the SARS-CoV-2 infection in large numbers during this pandemic, whereas others have shown a resilient side by manifesting only milder or no symptoms at all. This observation has relayed the onus of the heterogeneous pattern of SARS-CoV-2-induced critical illness among different populations to the host genetic factors. Here, the evolutionary route was explored and three genetic loci, i.e., rs10735079, rs2109069, and rs2236757, associated with COVID-19 were analyzed. Among the three, the risk allele A at genetic locus rs2236757 residing in the IFNAR2 gene was observed to have undergone recent positive selection in the African population.
Keywords: COVID-19; SNP variants; host genetic factors; natural selection; population genomics.
Copyright © 2022 Raza and Abbasi.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Study on the correlation between DPP9 rs2109069 and IFNAR2 rs2236757 polymorphisms with COVID-19 mortality.Nucleosides Nucleotides Nucleic Acids. 2025;44(1):41-56. doi: 10.1080/15257770.2024.2344179. Epub 2024 Apr 25. Nucleosides Nucleotides Nucleic Acids. 2025. PMID: 38660988
-
Association of IFNAR2 rs2236757 and OAS3 rs10735079 Polymorphisms with Susceptibility to COVID-19 Infection and Severity in Palestine.Interdiscip Perspect Infect Dis. 2023 Sep 16;2023:9551163. doi: 10.1155/2023/9551163. eCollection 2023. Interdiscip Perspect Infect Dis. 2023. PMID: 37745867 Free PMC article.
-
Genomic insight into COVID-19 severity in MAFLD patients: a single-center prospective cohort study.Front Genet. 2024 Sep 4;15:1460318. doi: 10.3389/fgene.2024.1460318. eCollection 2024. Front Genet. 2024. PMID: 39296547 Free PMC article.
-
COVID-19 pandemic: Insights into genetic susceptibility to SARS-CoV-2 and host genes implications on virus spread, disease severity and outcomes.Hum Antibodies. 2022;30(1):1-14. doi: 10.3233/HAB-211506. Hum Antibodies. 2022. PMID: 34864654 Review.
-
Immune response to SARS-CoV-2 variants: A focus on severity, susceptibility, and preexisting immunity.J Infect Public Health. 2022 Feb;15(2):277-288. doi: 10.1016/j.jiph.2022.01.007. Epub 2022 Jan 13. J Infect Public Health. 2022. PMID: 35074728 Free PMC article. Review.
Cited by
-
Participation of Single-Nucleotide Variants in IFNAR1 and IFNAR2 in the Immune Response against SARS-CoV-2 Infection: A Systematic Review.Pathogens. 2023 Nov 6;12(11):1320. doi: 10.3390/pathogens12111320. Pathogens. 2023. PMID: 38003785 Free PMC article. Review.
-
The Relationship between COVID-19 Severity in Children and Immunoregulatory Gene Polymorphism.Viruses. 2023 Oct 14;15(10):2093. doi: 10.3390/v15102093. Viruses. 2023. PMID: 37896870 Free PMC article.
-
Next-generation sequencing of host genetics risk factors associated with COVID-19 severity and long-COVID in Colombian population.Sci Rep. 2024 Apr 11;14(1):8497. doi: 10.1038/s41598-024-57982-3. Sci Rep. 2024. PMID: 38605121 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

