Development and characterization of a glycine biosensor system for fine-tuned metabolic regulation in Escherichia coli
- PMID: 35392910
- PMCID: PMC8991567
- DOI: 10.1186/s12934-022-01779-4
Development and characterization of a glycine biosensor system for fine-tuned metabolic regulation in Escherichia coli
Abstract
Background: In vivo biosensors have a wide range of applications, ranging from the detection of metabolites to the regulation of metabolic networks, providing versatile tools for synthetic biology and metabolic engineering. However, in view of the vast array of metabolite molecules, the existing number and performance of biosensors is far from sufficient, limiting their potential applications in metabolic engineering. Therefore, we developed the synthetic glycine-ON and -OFF riboswitches for metabolic regulation and directed evolution of enzyme in Escherichia coli.
Results: The results showed that a synthetic glycine-OFF riboswitch (glyOFF6) and an increased-detection-range synthetic glycine-ON riboswitch (glyON14) were successfully screened from a library based on the Bacillus subtilis glycine riboswitch using fluorescence-activated cell sorting (FACS) and tetA-based dual genetic selection. The two synthetic glycine riboswitches were successfully used in tunable regulation of lactate synthesis, dynamic regulation of serine synthesis and directed evolution of alanine-glyoxylate aminotransferase in Escherichia coli, respectively. Mutants AGXT22 and AGXT26 of alanine-glyoxylate aminotransferase with an increase of 58% and 73% enzyme activity were obtained by using a high-throughput screening platform based on the synthetic glycine-OFF riboswitch, and successfully used to increase the 5-aminolevulinic acid yield of engineered Escherichia coli.
Conclusions: A synthetic glycine-OFF riboswitch and an increased-detection-range synthetic glycine-ON riboswitch were successfully designed and screened. The developed riboswitches showed broad application in tunable regulation, dynamic regulation and directed evolution of enzyme in E. coli.
Keywords: Biosensor; Directed evolution; Escherichia coli; Glycine riboswitch; Metabolic regulation.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing financial interest.
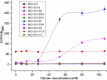
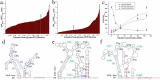
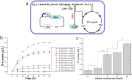
Figures




Similar articles
-
Characterization and Engineering of a Clostridium Glycine Riboswitch and Its Use To Control a Novel Metabolic Pathway for 5-Aminolevulinic Acid Production in Escherichia coli.ACS Synth Biol. 2019 Oct 18;8(10):2327-2335. doi: 10.1021/acssynbio.9b00137. Epub 2019 Oct 10. ACS Synth Biol. 2019. PMID: 31550137
-
Naringenin-responsive riboswitch-based fluorescent biosensor module for Escherichia coli co-cultures.Biotechnol Bioeng. 2017 Oct;114(10):2235-2244. doi: 10.1002/bit.26340. Epub 2017 Jun 5. Biotechnol Bioeng. 2017. PMID: 28543037
-
Artificial Caprolactam-Specific Riboswitch as an Intracellular Metabolite Sensor.ACS Synth Biol. 2019 Jun 21;8(6):1276-1283. doi: 10.1021/acssynbio.8b00452. Epub 2019 May 22. ACS Synth Biol. 2019. PMID: 31074964
-
Engineering and In Vivo Applications of Riboswitches.Annu Rev Biochem. 2017 Jun 20;86:515-539. doi: 10.1146/annurev-biochem-060815-014628. Epub 2017 Mar 30. Annu Rev Biochem. 2017. PMID: 28375743 Review.
-
Transcription-Factor-based Biosensor Engineering for Applications in Synthetic Biology.ACS Synth Biol. 2021 May 21;10(5):911-922. doi: 10.1021/acssynbio.0c00252. Epub 2021 Apr 25. ACS Synth Biol. 2021. PMID: 33899477 Review.
Cited by
-
Living in mangroves: a syntrophic scenario unveiling a resourceful microbiome.BMC Microbiol. 2024 Jun 28;24(1):228. doi: 10.1186/s12866-024-03390-6. BMC Microbiol. 2024. PMID: 38943070 Free PMC article.
-
Regulatory RNAs in Bacillus subtilis: A review on regulatory mechanism and applications in synthetic biology.Synth Syst Biotechnol. 2024 Feb 10;9(2):223-233. doi: 10.1016/j.synbio.2024.01.013. eCollection 2024 Jun. Synth Syst Biotechnol. 2024. PMID: 38385150 Free PMC article. Review.
-
Riboswitches, from cognition to transformation.Synth Syst Biotechnol. 2023 Jun 3;8(3):357-370. doi: 10.1016/j.synbio.2023.05.008. eCollection 2023 Sep. Synth Syst Biotechnol. 2023. PMID: 37325181 Free PMC article. Review.
-
Exploring L-isoleucine riboswitches for enhancing 4-hydroxyisoleucine production in Corynebacterium glutamicum.Biotechnol Lett. 2023 Sep;45(9):1169-1181. doi: 10.1007/s10529-023-03407-6. Epub 2023 Jul 3. Biotechnol Lett. 2023. PMID: 37395871
-
Development of a lysine biosensor for the dynamic regulation of cadaverine biosynthesis in E. coli.Microb Cell Fact. 2025 Jun 21;24(1):141. doi: 10.1186/s12934-025-02772-3. Microb Cell Fact. 2025. PMID: 40544235 Free PMC article.
References
-
- Gallagher RR, Li Z, Lewis AO, Isaacs FJ. Rapid editing and evolution of bacterial genomes using libraries of synthetic DNA. Nat Protoc. 2014;9(10):2301–2316. - PubMed
-
- Tong Y, Weber T, Lee SY. CRISPR/Cas-based genome engineering in natural product discovery. Nat Prod Rep. 2019;36(9):1262–1280. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

