Notch-dependent DNA cis-regulatory elements and their dose-dependent control of C. elegans stem cell self-renewal
- PMID: 35394007
- PMCID: PMC9058496
- DOI: 10.1242/dev.200332
Notch-dependent DNA cis-regulatory elements and their dose-dependent control of C. elegans stem cell self-renewal
Abstract
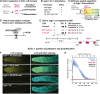
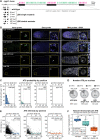
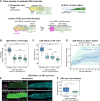
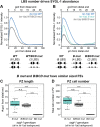
A long-standing biological question is how DNA cis-regulatory elements shape transcriptional patterns during metazoan development. Reporter constructs, cell culture assays and computational modeling have made major contributions to answering this question, but analysis of elements in their natural context is an important complement. Here, we mutate Notch-dependent LAG-1 binding sites (LBSs) in the endogenous Caenorhabditis elegans sygl-1 gene, which encodes a key stem cell regulator, and analyze the consequences on sygl-1 expression (nascent transcripts, mRNA, protein) and stem cell maintenance. Mutation of one LBS in a three-element cluster approximately halved both expression and stem cell pool size, whereas mutation of two LBSs essentially abolished them. Heterozygous LBS mutant clusters provided intermediate values. Our results lead to two major conclusions. First, both LBS number and configuration impact cluster activity: LBSs act additively in trans and synergistically in cis. Second, the SYGL-1 gradient promotes self-renewal above its functional threshold and triggers differentiation below the threshold. Our approach of coupling CRISPR/Cas9 LBS mutations with effects on both molecular and biological readouts establishes a powerful model for in vivo analyses of DNA cis-regulatory elements.
Keywords: sygl-1; Gradient; Homotypic cluster; Spatiotemporal resolution; Transcription factor binding site; smFISH.
© 2022. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interests The authors declare no competing or financial interests.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

