Mesenchymal tumor cells drive adaptive resistance of Trp53-/- breast tumor cells to inactivated mutant Kras
- PMID: 35398967
- PMCID: PMC9441006
- DOI: 10.1002/1878-0261.13220
Mesenchymal tumor cells drive adaptive resistance of Trp53-/- breast tumor cells to inactivated mutant Kras
Abstract
As precision medicine increases the response rate of treatment, tumors frequently bypass inhibition, and reoccur. In order for treatment to be effective long term, the mechanisms enabling treatment adaptation need to be understood. Here, we report a mouse model that, in the absence of p53 and the presence of oncogenic KrasG12D , develops breast tumors. Upon inactivation of KrasG12D , tumors initially regress and enter remission. Subsequently, the majority of tumors adapt to the withdrawal of KrasG12D expression and return. KrasG12D -independent tumor cells show a strong mesenchymal profile with active RAS-RAF-MEK-ERK (MAPK/ERK) signaling. Both KrasG12D -dependent and KrasG12D -independent tumors display a high level of genomic instability, and KrasG12D -independent tumors harbor numerous amplified genes that can activate the MAPK/ERK signaling pathway. Our study identifies both epithelial-mesenchymal transition (EMT) and active MAPK/ERK signaling in tumors that adapt to oncogenic KrasG12D withdrawal in a novel Trp53-/- breast cancer mouse model. To achieve long-lasting responses in the clinic to RAS-fueled cancer, treatment will need to focus in parallel on obstructing tumors from adapting to oncogene inhibition.
Keywords: Kras; breast cancer; mouse model; treatment adaptation.
© 2022 The Authors. Molecular Oncology published by John Wiley & Sons Ltd on behalf of Federation of European Biochemical Societies.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



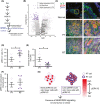

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

