LINC00511 enhances LUAD malignancy by upregulating GCNT3 via miR-195-5p
- PMID: 35399076
- PMCID: PMC8994914
- DOI: 10.1186/s12885-022-09459-7
LINC00511 enhances LUAD malignancy by upregulating GCNT3 via miR-195-5p
Abstract
Background: Accumulating evidence suggests that LINC00511 acts as an oncogenic long non-coding RNA (lncRNA) in various cancers, including lung adenocarcinoma (LUAD). Hence, we attempted to elucidate the potential role of LINC00511 in LUAD.
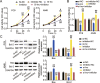
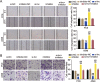
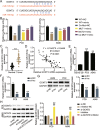
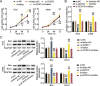
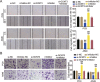
Methods: LINC00511, miR-195-5p, and GCNT3 expression in LUAD was detected by qRT-PCR. Changes in the proliferation, migration, and invasion of LUAD cells after abnormal regulation of LINC00511, miR-195-5p, or GCNT3 were detected by CCK-8, BrdU, wound healing, and transwell assays. Bax and Bcl-2 protein expression was measured by western blotting. Additionally, we identified the targeting effects of LINC00511, miR-195-5p, and GCNT3 using luciferase and RNA immunoprecipitation (RIP) assays.
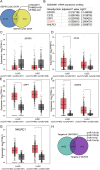
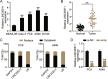
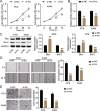
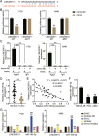
Results: LINC00511 and GCNT3 were found to be upregulated in LUAD, while miR-195-5p was downregulated. Silencing LINC00511 or GCNT3 decreased the proliferation, migration, invasion, and Bcl-2 protein content in LUAD cells and increased the expression of Bax. Interference with miR-195-5p promoted malignant proliferation of cancer cells. miR-195-5p expression was affected by LINC00511and targeted GCNT3.
Conclusion: Silencing LINC00511 promotes GCNT3 expression by inhibiting miR-195-5p and ultimately stimulates the malignant progression of LUAD.
Keywords: GCNT3; LINC00511; LUAD; miR-195-5p.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that there is no conflict of interests.
Figures
Similar articles
-
LncRNA LINC00511 promotes COL1A1-mediated proliferation and metastasis by sponging miR-126-5p/miR-218-5p in lung adenocarcinoma.BMC Pulm Med. 2022 Jul 16;22(1):272. doi: 10.1186/s12890-022-02070-3. BMC Pulm Med. 2022. PMID: 35842617 Free PMC article.
-
LINC00511 aggravates the malignancy of lung adenocarcinoma through sponging microRNA miR-4739 to regulate pyrroline-5-carboxylate reductase 1 expression.J Clin Lab Anal. 2022 Dec;36(12):e24760. doi: 10.1002/jcla.24760. Epub 2022 Nov 13. J Clin Lab Anal. 2022. PMID: 36371775 Free PMC article.
-
LINC02535/miR-30a-5p/GALNT3 axis contributes to lung adenocarcinoma progression via the NF- κ B signaling pathway.Cell Cycle. 2022 Dec;21(23):2455-2470. doi: 10.1080/15384101.2022.2101336. Epub 2022 Jul 19. Cell Cycle. 2022. PMID: 35852407 Free PMC article.
-
GCNT3-mediated glycosylation in cancer biology: Implications for tumorigenesis, metastasis, and therapeutic targeting.Int J Biol Macromol. 2025 Jun;315(Pt 2):144427. doi: 10.1016/j.ijbiomac.2025.144427. Epub 2025 May 20. Int J Biol Macromol. 2025. PMID: 40403799 Review.
-
The emerging roles of LINC00511 in breast cancer development and therapy.Front Oncol. 2024 Aug 14;14:1429262. doi: 10.3389/fonc.2024.1429262. eCollection 2024. Front Oncol. 2024. PMID: 39206156 Free PMC article. Review.
Cited by
-
Dendritic Cell-Related Immune Marker CD1C for Predicting Prognosis and Immunotherapy Opportunities of Lung Adenocarcinoma Patients.Appl Biochem Biotechnol. 2024 Dec;196(12):8724-8740. doi: 10.1007/s12010-024-04973-9. Epub 2024 Jun 22. Appl Biochem Biotechnol. 2024. PMID: 38907868
-
Feedback loop LINC00511-YTHDF2-SOX2 regulatory network drives cholangiocarcinoma progression and stemness.MedComm (2020). 2024 Oct 22;5(11):e743. doi: 10.1002/mco2.743. eCollection 2024 Nov. MedComm (2020). 2024. PMID: 39445001 Free PMC article.
-
A review on the role of LINC00511 in cancer.Front Genet. 2023 Apr 14;14:1116445. doi: 10.3389/fgene.2023.1116445. eCollection 2023. Front Genet. 2023. PMID: 37124625 Free PMC article. Review.
-
STLBRF: an improved random forest algorithm based on standardized-threshold for feature screening of gene expression data.Brief Funct Genomics. 2025 Jan 15;24:elae048. doi: 10.1093/bfgp/elae048. Brief Funct Genomics. 2025. PMID: 39736135 Free PMC article.
-
The Mutual Relationship between Glycosylation and Non-Coding RNAs in Cancer and Other Physio-Pathological Conditions.Int J Mol Sci. 2022 Dec 13;23(24):15804. doi: 10.3390/ijms232415804. Int J Mol Sci. 2022. PMID: 36555445 Free PMC article. Review.
References
-
- Travis WD, Brambilla E, Noguchi M, Nicholson AG, Geisinger K, Yatabe Y, et al. International Association for the Study of Lung Cancer/American Thoracic Society/European Respiratory Society: international multidisciplinary classification of lung adenocarcinoma: executive summary. Proc Am Thorac Soc. 2011;8(5):381–385. doi: 10.1513/pats.201107-042ST. - DOI - PubMed
-
- Costa GJ, de Mello MJG, Ferreira CG, Bergmann A, Thuler LCS. Increased incidence, morbidity and mortality rates for lung cancer in women in Brazil between 2000 and 2014: An analysis of three types of sources of secondary data. Lung Cancer. 2018;125:77–85. doi: 10.1016/j.lungcan.2018.09.005. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

