Differential Transcriptomics Analysis of IPEC-J2 Cells Single or Coinfected With Porcine Epidemic Diarrhea Virus and Transmissible Gastroenteritis Virus
- PMID: 35401515
- PMCID: PMC8989846
- DOI: 10.3389/fimmu.2022.844657
Differential Transcriptomics Analysis of IPEC-J2 Cells Single or Coinfected With Porcine Epidemic Diarrhea Virus and Transmissible Gastroenteritis Virus
Abstract
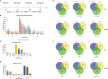
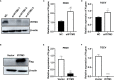
Porcine epidemic diarrhea (PED) and transmissible gastroenteritis (TGE) caused by porcine epidemic diarrhea virus (PEDV) and transmissible gastroenteritis virus (TGEV) are two highly contagious intestinal diseases in the swine industry worldwide. Notably, coinfection of TGEV and PEDV is common in piglets with diarrhea-related diseases. In this study, intestinal porcine epithelial cells (IPEC-J2) were single or coinfected with PEDV and/or TGEV, followed by the comparison of differentially expressed genes (DEGs), especially interferon-stimulated genes (ISGs), between different groups via transcriptomics analysis and real-time qPCR. The antiviral activity of swine interferon-induced transmembrane protein 3 (sIFITM3) on PEDV and TGEV infection was also evaluated. The results showed that DEGs can be detected in the cells infected with PEDV, TGEV, and PEDV+TGEV at 12, 24, and 48 hpi, and the number of DEGs was the highest at 24 hpi. The DEGs are mainly annotated to the GO terms of protein binding, immune system process, organelle part, and intracellular organelle part. Furthermore, 90 ISGs were upregulated during PEDV or TGEV infection, 27 of which were associated with antiviral activity, including ISG15, OASL, IFITM1, and IFITM3. Furthermore, sIFITM3 can significantly inhibit PEDV and TGEV infection in porcine IPEC-J2 cells and/or monkey Vero cells. Besides, sIFITM3 can also inhibit vesicular stomatitis virus (VSV) replication in Vero cells. These results indicate that sIFITM3 has broad-spectrum antiviral activity.
Keywords: coinfection; differential transcriptomics; interferon-induced transmembrane protein (IFITM); interferon-stimulated genes (ISGs); porcine epidemic diarrhea virus (PEDV); transmissible gastroenteritis virus (TGEV).
Copyright © 2022 Song, Chen, Hao, Jiang, Xu, Li, Chen, Gao, Jin, Ren and Li.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures







Similar articles
-
Surface-Displayed Porcine IFN-λ3 in Lactobacillus plantarum Inhibits Porcine Enteric Coronavirus Infection of Porcine Intestinal Epithelial Cells.J Microbiol Biotechnol. 2020 Apr 28;30(4):515-525. doi: 10.4014/jmb.1909.09041. J Microbiol Biotechnol. 2020. PMID: 31838830 Free PMC article.
-
Decreased NHE3 activity in intestinal epithelial cells in TGEV and PEDV-induced piglet diarrhea.Vet Microbiol. 2021 Dec;263:109263. doi: 10.1016/j.vetmic.2021.109263. Epub 2021 Oct 27. Vet Microbiol. 2021. PMID: 34749283
-
Establishment and application of a quadruplex real-time RT-qPCR assay for differentiation of TGEV, PEDV, PDCoV, and PoRVA.Microb Pathog. 2024 Jun;191:106646. doi: 10.1016/j.micpath.2024.106646. Epub 2024 Apr 16. Microb Pathog. 2024. PMID: 38631414
-
Immune evasion of porcine enteric coronaviruses and viral modulation of antiviral innate signaling.Virus Res. 2016 Dec 2;226:128-141. doi: 10.1016/j.virusres.2016.05.015. Epub 2016 May 19. Virus Res. 2016. PMID: 27212682 Free PMC article. Review.
-
Advances research in porcine enteric coronavirus therapies and antiviral drugs.Vet Q. 2024 Dec;44(1):1-49. doi: 10.1080/01652176.2024.2421299. Epub 2024 Nov 1. Vet Q. 2024. PMID: 39484691 Free PMC article. Review.
Cited by
-
Chicken interferon-induced transmembrane proteins inhibit Newcastle disease virus infection by affecting viral entry and W protein expression.Vet Res. 2025 May 21;56(1):104. doi: 10.1186/s13567-025-01530-y. Vet Res. 2025. PMID: 40399912 Free PMC article.
-
Differences in cytokines expression between Vero cells and IPEC-J2 cells infected with porcine epidemic diarrhea virus.Front Microbiol. 2022 Nov 10;13:1002349. doi: 10.3389/fmicb.2022.1002349. eCollection 2022. Front Microbiol. 2022. PMID: 36439802 Free PMC article.
-
Antiviral Effect of pIFNLs against PEDV and VSV Infection in Different Cells.Int J Mol Sci. 2022 Aug 26;23(17):9661. doi: 10.3390/ijms23179661. Int J Mol Sci. 2022. PMID: 36077060 Free PMC article.
-
NS7a of SADS-CoV promotes viral infection via inducing apoptosis to suppress type III interferon production.J Virol. 2024 May 14;98(5):e0031724. doi: 10.1128/jvi.00317-24. Epub 2024 Apr 16. J Virol. 2024. PMID: 38624231 Free PMC article.
-
Bivalent circular RNA vaccines against porcine epidemic diarrhea virus and transmissible gastroenteritis virus.Front Immunol. 2025 Mar 31;16:1562865. doi: 10.3389/fimmu.2025.1562865. eCollection 2025. Front Immunol. 2025. PMID: 40230842 Free PMC article.
References
-
- Zhu Y, Liang L, Luo Y, Wang G, Wang C, Cui Y, et al. . A Sensitive Duplex Nanoparticle-Assisted PCR Assay for Identifying Porcine Epidemic Diarrhea Virus and Porcine Transmissible Gastroenteritis Virus From Clinical Specimens. Virus Genes (2017) 53:71–6. doi: 10.1007/s11262-016-1405-z - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

