Flagella at the Host-Microbe Interface: Key Functions Intersect With Redundant Responses
- PMID: 35401545
- PMCID: PMC8987104
- DOI: 10.3389/fimmu.2022.828758
Flagella at the Host-Microbe Interface: Key Functions Intersect With Redundant Responses
Abstract
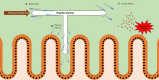
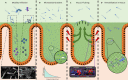
Many bacteria and other microbes achieve locomotion via flagella, which are organelles that function as a swimming motor. Depending on the environment, flagellar motility can serve a variety of beneficial functions and confer a fitness advantage. For example, within a mammalian host, flagellar motility can provide bacteria the ability to resist clearance by flow, facilitate access to host epithelial cells, and enable travel to nutrient niches. From the host's perspective, the mobility that flagella impart to bacteria can be associated with harmful activities that can disrupt homeostasis, such as invasion of epithelial cells, translocation across epithelial barriers, and biofilm formation, which ultimately can decrease a host's reproductive fitness from a perspective of natural selection. Thus, over an evolutionary timescale, the host developed a repertoire of innate and adaptive immune countermeasures that target and mitigate this microbial threat. These countermeasures are wide-ranging and include structural components of the mucosa that maintain spatial segregation of bacteria from the epithelium, mechanisms of molecular recognition and inducible responses to flagellin, and secreted effector molecules of the innate and adaptive immune systems that directly inhibit flagellar motility. While much of our understanding of the dynamics of host-microbe interaction regarding flagella is derived from studies of enteric bacterial pathogens where flagella are a recognized virulence factor, newer studies have delved into host interaction with flagellated members of the commensal microbiota during homeostasis. Even though many aspects of flagellar motility may seem innocuous, the host's redundant efforts to stop bacteria in their tracks highlights the importance of this host-microbe interaction.
Keywords: DEFA6; IBD; IgA; LYPD8; TLR5; ZG16; fliC; goblet cell.
Copyright © 2022 Akahoshi and Bevins.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

