First Principles Calculation of Protein-Protein Dimer Affinities of ALS-Associated SOD1 Mutants
- PMID: 35402516
- PMCID: PMC8988244
- DOI: 10.3389/fmolb.2022.845013
First Principles Calculation of Protein-Protein Dimer Affinities of ALS-Associated SOD1 Mutants
Abstract
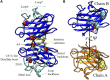
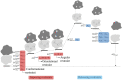
Cu,Zn superoxide dismutase (SOD1) is a 32 kDa homodimer that converts toxic oxygen radicals in neurons to less harmful species. The dimerization of SOD1 is essential to the stability of the protein. Monomerization increases the likelihood of SOD1 misfolding into conformations associated with aggregation, cellular toxicity, and neuronal death in familial amyotrophic lateral sclerosis (fALS). The ubiquity of disease-associated mutations throughout the primary sequence of SOD1 suggests an important role of physicochemical processes, including monomerization of SOD1, in the pathology of the disease. Herein, we use a first-principles statistical mechanics method to systematically calculate the free energy of dimer binding for SOD1 using molecular dynamics, which involves sequentially computing conformational, orientational, and separation distance contributions to the binding free energy. We consider the effects of two ALS-associated mutations in SOD1 protein on dimer stability, A4V and D101N, as well as the role of metal binding and disulfide bond formation. We find that the penalty for dimer formation arising from the conformational entropy of disordered loops in SOD1 is significantly larger than that for other protein-protein interactions previously considered. In the case of the disulfide-reduced protein, this leads to a bound complex whose formation is energetically disfavored. Somewhat surprisingly, the loop free energy penalty upon dimerization is still significant for the holoprotein, despite the increased structural order induced by the bound metal cations. This resulted in a surprisingly modest increase in dimer binding free energy of only about 1.5 kcal/mol upon metalation of the protein, suggesting that the most significant stabilizing effects of metalation are on folding stability rather than dimer binding stability. The mutant A4V has an unstable dimer due to weakened monomer-monomer interactions, which are manifested in the calculation by a separation free energy surface with a lower barrier. The mutant D101N has a stable dimer partially due to an unusually rigid β-barrel in the free monomer. D101N also exhibits anticooperativity in loop folding upon dimerization. These computational calculations are, to our knowledge, the most quantitatively accurate calculations of dimer binding stability in SOD1 to date.
Keywords: amyotrophic lateral sclerosis; dimer dissociation; free energy perturbation; loop entropy; molecular dynamics simulations; protein misfolding (conformational) diseases; protein–protein interactions; superoxide dismutase (Cu–Zn).
Copyright © 2022 Hsueh, Nijland, Peng, Hilton and Plotkin.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
-
- Abdolvahabi A., Shi Y., Rasouli S., Croom C. M., Aliyan A., Martí A. A., et al. (2017). Kaplan-meier Meets Chemical Kinetics: Intrinsic Rate of SOD1 Amyloidogenesis Decreased by Subset of ALS Mutations and Cannot Fully Explain Age of Disease Onset. ACS Chem. Neurosci. 8 (6), 1378–1389. 10.1021/acschemneuro.7b00029 - DOI - PubMed
-
- Abraham M. J., Murtola T., Schulz R., Páll S., Smith J. C., Hess B., et al. (2015). GROMACS: High Performance Molecular Simulations through Multi-Level Parallelism from Laptops to Supercomputers. SoftwareX 1, 19–25. 10.1016/j.softx.2015.06.001 - DOI
LinkOut - more resources
Full Text Sources
Miscellaneous

