Entamoeba histolytica EHD1 Is Involved in Mitosome-Endosome Contact
- PMID: 35404118
- PMCID: PMC9040739
- DOI: 10.1128/mbio.03849-21
Entamoeba histolytica EHD1 Is Involved in Mitosome-Endosome Contact
Abstract
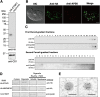
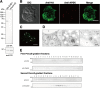
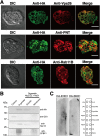
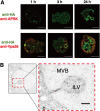
Interorganellar cross talk is often mediated by membrane contact sites (MCSs), which are zones where participating membranes come within 30 nm of one another. MCSs have been found in organelles, including the endoplasmic reticulum, Golgi bodies, endosomes, and mitochondria. Despite its seeming ubiquity, reports of MCS involving mitochondrion-related organelles (MROs) present in a few anaerobic parasitic protozoa remain lacking. Entamoeba histolytica, the etiological agent of amoebiasis, possesses an MRO called the mitosome. We previously discovered several Entamoeba-specific transmembrane mitosomal proteins (ETMPs) from in silico and cell-biological analyses. One of them, ETMP1 (EHI_175060), was predicted to have one transmembrane domain and two coiled-coil regions and was demonstrated to be mitosome membrane integrated based on carbonate fractionation and immunoelectron microscopy (IEM) data. Immunoprecipitation analysis detected a candidate interacting partner, EH domain-containing protein (EHD1; EHI_105270). We expressed hemagglutinin (HA)-tagged EHD1 in E. histolytica, and subsequent immunofluorescence and IEM data indicated an unprecedented MCS between the mitosome and the endosome. Live imaging of a green fluorescent protein (GFP)-EHD1-expressing strain demonstrated that EHD1 is involved in early endosome formation and is observed in MCS between endosomes of various sizes. In vitro assays using recombinant His-EHD1 demonstrated ATPase activity. MCSs are involved in lipid transfer, ion homeostasis, and organelle dynamics. The serendipitous discovery of the ETMP1-interacting partner EHD1 led to the observation of the mitosome-endosome contact site in E. histolytica. It opened a new view of how the relic mitochondria of Entamoeba may likewise be involved in organelle cross talk, a conserved feature of mitochondria and other organelles in general. IMPORTANCE Membrane contact sites (MCSs) are key regulators of interorganellar communication and have been widely demonstrated between various organelles. However, studies on MCSs involving mitochondrion-related organelles (MROs), present in some anaerobic parasitic protozoans, remain scarce. Entamoeba histolytica, the etiological agent of amoebiasis, possesses an MRO called the mitosome. This organelle is crucial for cellular differentiation and disease transmission, thereby significantly contributing to the amoeba's parasitic lifestyle. Our recent discovery of the interaction between the Entamoeba-specific transmembrane mitosomal protein (ETMP1) and EH domain-containing protein (EHD1) showcases a newly found mitosome-endosome contact site in E. histolytica. This finding reflects the idea that despite their substantially divergent and reduced nature, MROs like mitosomes conserve mechanisms for interorganellar cross talk. We posit lipid and ion transport, mitosome fission, and quality control as potential processes that are mediated by the ETMP1-EHD1-tethered mitosome-endosome contact site in E. histolytica.
Keywords: EH domain; Entamoeba histolytica; endosome; membrane contact site; mitochondrion-related organelles; mitosome.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






Similar articles
-
The mitosome of the anaerobic parasitic protist Entamoeba histolytica: A peculiar and minimalist mitochondrion-related organelle.J Eukaryot Microbiol. 2022 Nov;69(6):e12923. doi: 10.1111/jeu.12923. Epub 2022 Jun 4. J Eukaryot Microbiol. 2022. PMID: 35588086 Free PMC article. Review.
-
An Entamoeba-Specific Mitosomal Membrane Protein with Potential Association to the Golgi Apparatus.Genes (Basel). 2019 May 13;10(5):367. doi: 10.3390/genes10050367. Genes (Basel). 2019. PMID: 31086122 Free PMC article.
-
Screening and discovery of lineage-specific mitosomal membrane proteins in Entamoeba histolytica.Mol Biochem Parasitol. 2016 Sep-Oct;209(1-2):10-17. doi: 10.1016/j.molbiopara.2016.01.001. Epub 2016 Jan 11. Mol Biochem Parasitol. 2016. PMID: 26792249
-
Interorganellar communication and membrane contact sites in protozoan parasites.Parasitol Int. 2021 Aug;83:102372. doi: 10.1016/j.parint.2021.102372. Epub 2021 Apr 29. Parasitol Int. 2021. PMID: 33933652 Review.
-
Import of Entamoeba histolytica Mitosomal ATP Sulfurylase Relies on Internal Targeting Sequences.Microorganisms. 2020 Aug 12;8(8):1229. doi: 10.3390/microorganisms8081229. Microorganisms. 2020. PMID: 32806678 Free PMC article.
Cited by
-
Endocytosis in anaerobic parasitic protists.Mem Inst Oswaldo Cruz. 2024 Jul 26;119:e240058. doi: 10.1590/0074-02760240058. eCollection 2024. Mem Inst Oswaldo Cruz. 2024. PMID: 39082582 Free PMC article. Review.
-
Encystation and Stress Responses under the Control of Ubiquitin-like Proteins in Pathogenic Amoebae.Microorganisms. 2023 Oct 31;11(11):2670. doi: 10.3390/microorganisms11112670. Microorganisms. 2023. PMID: 38004682 Free PMC article. Review.
-
The mitosome of the anaerobic parasitic protist Entamoeba histolytica: A peculiar and minimalist mitochondrion-related organelle.J Eukaryot Microbiol. 2022 Nov;69(6):e12923. doi: 10.1111/jeu.12923. Epub 2022 Jun 4. J Eukaryot Microbiol. 2022. PMID: 35588086 Free PMC article. Review.
-
A missense mutation in Ehd1 associated with defective spermatogenesis and male infertility.Front Cell Dev Biol. 2023 Oct 12;11:1240558. doi: 10.3389/fcell.2023.1240558. eCollection 2023. Front Cell Dev Biol. 2023. PMID: 37900275 Free PMC article.
-
Pathogenicity and virulence of Entamoeba histolytica, the agent of amoebiasis.Virulence. 2023 Dec;14(1):2158656. doi: 10.1080/21505594.2022.2158656. Virulence. 2023. PMID: 36519347 Free PMC article. Review.
References
-
- Scorrano L, De Matteis MA, Emr S, Giordano F, Hajnóczky G, Kornmann B, Lackner LL, Levine TP, Pellegrini L, Reinisch K, Rizzuto R, Simmen T, Stenmark H, Ungermann C, Schuldiner M. 2019. Coming together to define membrane contact sites. Nat Commun 10:1287. doi:10.1038/s41467-019-09253-3. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

