Modified eQTL and Somatic DNA Segment Alterations in Esophageal Squamous Cell Carcinoma for Genes Related to Immunity, DNA Repair, and Inflammation
- PMID: 35406404
- PMCID: PMC8996990
- DOI: 10.3390/cancers14071629
Modified eQTL and Somatic DNA Segment Alterations in Esophageal Squamous Cell Carcinoma for Genes Related to Immunity, DNA Repair, and Inflammation
Abstract
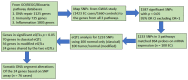
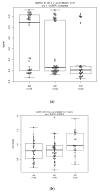
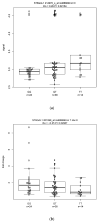
We integrated ESCC expression and GWAS genotyping, to investigate eQTL and somatic DNA segment alterations, including somatic copy number alteration, allelic imbalance (AI), and loss of heterozygosity (LOH) in ESCC. First, in eQTL analysis, we used a classical approach based on genotype data from GWAS and expression signals in normal tissue samples, and then used a modified approach based on fold change in the tumor vs. normal samples. We focused on the genes in three pathways: inflammation, DNA repair, and immunity. Among the significant (p < 0.05) SNP-probe pairs from classical and modified eQTL analyses, 24 genes were shared by the two approaches, including 18 genes that showed the same numbers of SNPs and probes and 6 genes that had the different numbers of SNPs and probes. For these 18 genes, we found 28 SNP−probe pairs were correlated in opposite directions in the two approaches, indicating an intriguing difference between the classical and modified eQTL approaches. Second, we analyzed the somatic DNA segment alterations. Across the 24 genes, abnormal gene expression on mRNA arrays was seen in 19−95% of cases and 26−78% showed somatic DNA segment alterations on Affymetrix GeneChip Human Mapping Arrays. The results suggested that this strategy could identify gene expression and somatic DNA segment alterations for biological markers (genes) by combining classical and modified eQTLs and somatic DNA evaluation on SNP arrays. Thus, this study approach may allow us to understand functionality indicative of potentially relevant biomarkers in ESCC.
Keywords: DNA segment alterations; ESCC; SNP; eQTL.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Li J.Y. Epidemiology of esophageal cancer in China. Natl. Cancer Inst. Monogr. 1982;62:113–120. - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

