Higher Phytohormone Contents and Weaker Phytohormone Signal Transduction Were Observed in Cold-Tolerant Cucumber
- PMID: 35406941
- PMCID: PMC9003209
- DOI: 10.3390/plants11070961
Higher Phytohormone Contents and Weaker Phytohormone Signal Transduction Were Observed in Cold-Tolerant Cucumber
Abstract
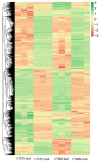
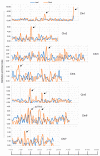
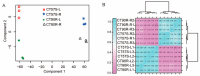
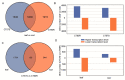
Cucumbers (Cucumis sativus L.) originated from the South Asian subcontinent, and most of them are fragile to cold stress. In this study, we evaluated the cold tolerance of 115 cucumber accessions and screened out 10 accessions showing high resistance to cold stress. We measured and compared plant hormone contents between cold-tolerant cucumber CT90R and cold-sensitive cucumber CT57S in cold treatment. Most of the detected plant hormones showed significantly higher content in CT90R. To elucidate the role of plant hormones, we compared the leaf- and root-transcriptomes of CT90R with those of CT57S in cold stress treatment. In leaves, there were 1209 differentially expressed genes (DEGs) between CT90R and CT57S, while there were 703 in roots. These DEGs were not evenly distributed across the chromosomes and there were significant enrichments at particular positions, including qLTT6.2, a known QTL controlling cucumber cold tolerance. The GO and KEGG enrichment analysis showed that there was a significant difference in the pathway of plant hormone transductions between CT90R and CT57S in leaves. In short, genes involved in plant hormone transductions showed lower transcription levels in CT90R. In roots, the most significantly different pathway was phenylpropanoid biosynthesis. CT90R seemed to actively accumulate more monolignols by upregulating cinnamyl-alcohol dehydrogenase (CAD) genes. These results above suggest a new perspective on the regulation mechanism of cold tolerance in cucumbers.
Keywords: Cucumis sativus; RNA-seq; cold tolerance; plant hormone; transcriptome.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
-
- Smeets L., Wehner T.C. Environmental effects on genetic variation of chilling resistance in cucumber. Euphytica. 1997;97:217–225. doi: 10.1023/A:1003084821178. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

