Generative Adversarial Networks for Creating Synthetic Nucleic Acid Sequences of Cat Genome
- PMID: 35409058
- PMCID: PMC8998662
- DOI: 10.3390/ijms23073701
Generative Adversarial Networks for Creating Synthetic Nucleic Acid Sequences of Cat Genome
Abstract
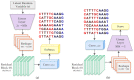
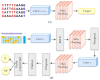
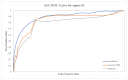
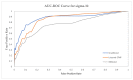
Nucleic acids are the basic units of deoxyribonucleic acid (DNA) sequencing. Every organism demonstrates different DNA sequences with specific nucleotides. It reveals the genetic information carried by a particular DNA segment. Nucleic acid sequencing expresses the evolutionary changes among organisms and revolutionizes disease diagnosis in animals. This paper proposes a generative adversarial networks (GAN) model to create synthetic nucleic acid sequences of the cat genome tuned to exhibit specific desired properties. We obtained the raw sequence data from Illumina next generation sequencing. Various data preprocessing steps were performed using Cutadapt and DADA2 tools. The processed data were fed to the GAN model that was designed following the architecture of Wasserstein GAN with gradient penalty (WGAN-GP). We introduced a predictor and an evaluator in our proposed GAN model to tune the synthetic sequences to acquire certain realistic properties. The predictor was built for extracting samples with a promoter sequence, and the evaluator was built for filtering samples that scored high for motif-matching. The filtered samples were then passed to the discriminator. We evaluated our model based on multiple metrics and demonstrated outputs for latent interpolation, latent complementation, and motif-matching. Evaluation results showed our proposed GAN model achieved 93.7% correlation with the original data and produced significant outcomes as compared to existing models for sequence generation.
Keywords: WGAN-GP; cat genome; generative adversarial networks; motif matching; nucleic acid sequences; promoter classification; promoter prediction; synthetic genome.
Conflict of interest statement
The authors declare no conflict of interest regarding the design of this study, analyses and writing of this manuscript.
Figures

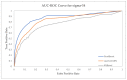



References
-
- Nouws S., Bogaerts B., Verhaegen B., Denayer S., Piérard D., Marchal K., Roosens N.H., Vanneste K., De Keersmaecker S.C. Impact of DNA extraction on whole genome sequencing analysis for characterization and relatedness of Shiga toxin-producing Escherichia coli isolates. Sci. Rep. 2020;10:14649. doi: 10.1038/s41598-020-71207-3. - DOI - PMC - PubMed
-
- Goodfellow I., Pouget-Abadie J., Mirza M., Xu B., Warde-Farley D., Ozair S., Courville A., Bengio Y. Generative adversarial nets. Adv. Neural Inf. Process. Syst. 2014;27 doi: 10.48550/ARXIV.1406.2661. - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

