Transcriptome Analysis of Populus euphratica under Salt Treatment and PeERF1 Gene Enhances Salt Tolerance in Transgenic Populus alba × Populus glandulosa
- PMID: 35409087
- PMCID: PMC8998595
- DOI: 10.3390/ijms23073727
Transcriptome Analysis of Populus euphratica under Salt Treatment and PeERF1 Gene Enhances Salt Tolerance in Transgenic Populus alba × Populus glandulosa
Abstract
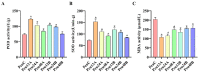
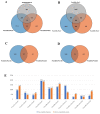
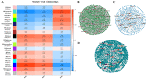
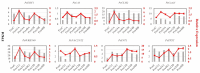
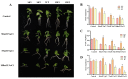
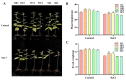
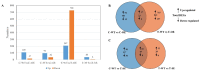
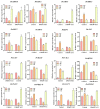
Populus euphratica is mainly distributed in desert environments with dry and hot climate in summer and cold in winter. Compared with other poplars, P. euphratica is more resistant to salt stress. It is critical to investigate the transcriptome and molecular basis of salt tolerance in order to uncover stress-related genes. In this study, salt-tolerant treatment of P. euphratica resulted in an increase in osmo-regulatory substances and recovery of antioxidant enzymes. To improve the mining efficiency of candidate genes, the analysis combining both the transcriptome WGCNA and the former GWAS results was selected, and a range of key regulatory factors with salt resistance were found. The PeERF1 gene was highly connected in the turquoise modules with significant differences in salt stress traits, and the expression levels were significantly different in each treatment. For further functional verification of PeERF1, we obtained stable overexpression and dominant suppression transgenic lines by transforming into Populus alba × Populusglandulosa. The growth and physiological characteristics of the PeERF1 overexpressed plants were better than that of the wild type under salt stress. Transcriptome analysis of leaves of transgenic lines and WT revealed that highly enriched GO terms in DEGs were associated with stress responses, including abiotic stimuli responses, chemical responses, and oxidative stress responses. The result is helpful for in-depth analysis of the salt tolerance mechanism of poplar. This work provides important genes for poplar breeding with salt tolerance.
Keywords: PeERF1; Populus euphratica; WGCNA; salt stress; transcriptome.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




References
-
- Kazuo S., Kazuko Y.S. Gene networks involved in drought stress response and tolerance. J. Exp. Bot. 2007;58:221–227. - PubMed
-
- Zeng F., Shabala S., Maksimović J.D., Maksimović V., Bonales-Alatorre E., Shabala L., Yu M., Zhang G., Živanović B.D. Revealing mechanisms of salinity tissue tolerance in succulent halophytes: A case study for Carpobrotus rossi. Plant Cell Environ. 2018;41:2654–2667. doi: 10.1111/pce.13391. - DOI - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

