Role and Regulation of Pif1 Family Helicases at the Replication Fork
- PMID: 35409096
- PMCID: PMC8998199
- DOI: 10.3390/ijms23073736
Role and Regulation of Pif1 Family Helicases at the Replication Fork
Abstract
Pif1 helicases are a multifunctional family of DNA helicases that are important for many aspects of genomic stability in the nucleus and mitochondria. Pif1 helicases are conserved from bacteria to humans. Pif1 helicases play multiple roles at the replication fork, including promoting replication through many barriers such as G-quadruplex DNA, the rDNA replication fork barrier, tRNA genes, and R-loops. Pif1 helicases also regulate telomerase and promote replication termination, Okazaki fragment maturation, and break-induced replication. This review highlights many of the roles and regulations of Pif1 at the replication fork that promote cellular health and viability.
Keywords: DNA helicase; G-quadruplexes; Pif1 helicase; replication fork; replication fork barriers; telomerase.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analysis, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures


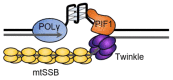
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

