Heterogeneity in extracellular vesicle secretion by single human macrophages revealed by super-resolution microscopy
- PMID: 35415881
- PMCID: PMC9006015
- DOI: 10.1002/jev2.12215
Heterogeneity in extracellular vesicle secretion by single human macrophages revealed by super-resolution microscopy
Abstract
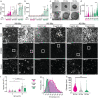
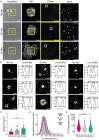
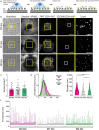
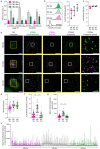
The diverse origins, nanometre-scale and invasive isolation procedures associated with extracellular vesicles (EVs) mean they are usually studied in bulk and disconnected from their parental cell. Here, we used super-resolution microscopy to directly compare EVs secreted by individual human monocyte-derived macrophages (MDMs). MDMs were differentiated to be M0-, M1- or M2-like, with all three secreting EVs at similar densities following activation. However, M0-like cells secreted larger EVs than M1- and M2-like macrophages. Proteomic analysis revealed variations in the contents of differently sized EVs as well as between EVs secreted by different MDM phenotypes. Super resolution microscopy of single-cell secretions identified that the class II MHC protein, HLA-DR, was expressed on ∼40% of EVs secreted from M1-like MDMs, which was double the frequency observed for M0-like and M2-like EVs. Strikingly, human macrophages, isolated from the resected lungs of cancer patients, secreted EVs that expressed HLA-DR at double the frequency and with greater intensity than M1-like EVs. Quantitative analysis of single-cell EV profiles from all four macrophage phenotypes revealed distinct secretion types, five of which were consistent across multiple sample cohorts. A sub-population of M1-like MDMs secreted EVs similar to lung macrophages, suggesting an expansion or recruitment of cells with a specific EV secretion profile within the lungs of cancer patients. Thus, quantitative analysis of EV heterogeneity can be used for single cell profiling and to reveal novel macrophage biology.
Keywords: class II MHC protein; extracellular vesicles; macrophages; single-cell analysis; super-resolution microscopy.
© 2022 The Authors. Journal of Extracellular Vesicles published by Wiley Periodicals, LLC on behalf of the International Society for Extracellular Vesicles.
Conflict of interest statement
G.M.T. is employed by G.S.K. and D.M.D. is a consultant and advisor to G.S.K.
Figures


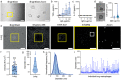

References
-
- Ambrose, A. R. , Dechantsreiter, S. , Shah, R. , Angeles Montero, M. , Quinn, A. M. , Hessel, E. M. , Beinke, S. , Tannahill, G. M. , & Davis, D. M. (2020).Corrected Super‐Resolution Microscopy Enables Nanoscale Imaging of Autofluorescent Lung Macrophages. Biophysical Journal, 119, 2403–2417. - PMC - PubMed
-
- Ambrose, A. R. , Hazime, K. S. , Worboys, J. D. , Niembro‐Vivanco, O. , & Davis, D. M. (2020). Synaptic secretion from human natural killer cells is diverse and includes supramolecular attack particles. Proceedings of the National Academy of Sciences of the United States of America, 117, 23717–23720. - PMC - PubMed
-
- Bertani, F. R. , Mozetic, P. , Fioramonti, M. , Iuliani, M. , Ribelli, G. , Pantano, F. , Santini, D. , Tonini, G. , Trombetta, M. , Businaro, L. , Selci, S. , & Rainer, A. (2017). Classification of M1/M2‐polarized human macrophages by label‐free hyperspectral reflectance confocal microscopy and multivariate analysis. Science Reports, 7, 1–9. - PMC - PubMed
-
- Brown, A. C. N. , Oddos, S. , Dobbie, I. M. , Alakoskela, J. M. , Parton, R. M. , Eissmann, P. , Neil, M. A. A. , Dunsby, C. , French, P. M. W. , Davis, I. , & Davis, D. M. (2011). Remodelling of cortical actin where lytic granules dock at natural killer cell immune synapses revealed by super‐resolution microscopy. PLOS Biology, 9(9), e1001152. - PMC - PubMed
-
- Buschow, S. I. , Hoen, E. N. M N‐'t , Niel, G. , Pols, M. S. , Broeke, T. , Lauwen, M. , Ossendorp, F. , Melief, C. J. M. , Raposo, G. , Wubbolts, R. , Wauben, M. H. M. , & Stoorvogel, W. (2009). MHC II in Dendritic Cells is Targeted to Lysosomes or T Cell‐Induced Exosomes Via Distinct Multivesicular Body Pathways. Traffic (Copenhagen, Denmark), 10, 1528–1542. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

